Cats and genes: 40 years later
There are half a billion domestic cats on Earth. They catch mice, feud with dogs and pigeons – and are a remarkably good object of genetic studies. Here, experts in genetics talk about the modern knowledge of the structure, functions and evolution of the cat genome and discuss the perspectives of cat genetics in the new decade
Pavel Borodin: in previous episodes
40 years ago, I published my first paper on cat genetics in “Chemisty and life”. I tried to persuade the reader that the cat, and not the pea or the fruitfly, is the best showcase of genetics.
Back then, cat genetics was basically limited to describing exotic phenotypes and the laws of their inheritance. Essentially, this was enough to create and sustain breeds and for genogeographic studies. Later we discovered that some of the hypotheses of that time about the inheritance of the simplest and most obvious traits were, to put it mildly, false. In most cases, the path from gene to trait was unknown; at best, it was tentatively defined in a mouse model and brazenly attributed to the cat. We knew that the cat genome contained DNA, but this was the limit of our knowledge of molecular genetics. Forty years ago, it was clear that the cat was related to the lion and the tiger, but the details of this relation were fogged by eons of evolution.
In 1989, I published another article in the same magazine, called “Cats and genes: ten years later”. The readers saw how the cat chromosomes looked under the microscope, learned the structure of the first genetic maps of this mammal and saw for themselves the incredible similarity of these maps to those of the human. I finished the article with a promise to write a sequel, called “Cats and genes: 20 years later”. I did not keep my promise.
However, I did write a book summing up the achievements of cat genetics in the XX century. It saw light in 1995 and has been reprinted countless times, including a reprint in the “Masterpieces of popular science” (sic!) series. The book was included in the long list of the Enlightener Award and got the Special Diploma of the Scientific Journalists Club, with a very true wording: ”For efficient use of cats to popularize science”.
 In 2009, “Science First Hand” published my article called “Cats and genes: 30 years later”. The 2000’s saw a boom in cat genetics. The cat genome was sequenced and partially annotated, and the first cloned and transgenic cats were born. Comparative genomics allowed to reconstruct the main stages of the mammalian evolution in general and the evolution of felids in particular. It turned out that the last common ancestor of the cat, the horse, and the bat lived relatively recently – around 79 million years ago. This is what I wrote about in the article. The last common ancestor story spread across the Runet under the title:“The cat and the horse have the same common ancestor, – Pavel Borodin”. This is how I became the ancestor of the cat and the horse.
In 2009, “Science First Hand” published my article called “Cats and genes: 30 years later”. The 2000’s saw a boom in cat genetics. The cat genome was sequenced and partially annotated, and the first cloned and transgenic cats were born. Comparative genomics allowed to reconstruct the main stages of the mammalian evolution in general and the evolution of felids in particular. It turned out that the last common ancestor of the cat, the horse, and the bat lived relatively recently – around 79 million years ago. This is what I wrote about in the article. The last common ancestor story spread across the Runet under the title:“The cat and the horse have the same common ancestor, – Pavel Borodin”. This is how I became the ancestor of the cat and the horse.
10 more years passed. It’s time for “Cats and genes: 40 years later”, and we are writing it together with Lyubov’ Malinovskaya. When Luybov’came to our lab, I asked her for help with the upcoming edition of Cats and Genes. And she did it – brilliantly! She gathered all freshly published materials on cat genetics, proofread everything and tracked out errors and misprints which had migrated from the previous articles. Shortly afterwards, she bought a male cat. A bit later, she bought a female cat. She then has passed the felinology course and registered her own cattery. I am thoroughly convinced that it was the work with my book that introduced her to the modest charm of cats and cat science, even though she has been denying this bluntly – let it be on her own head.
The important thing is, we wrote this article together. This is a foundation assuring that the tradition will live on, and that every decade, be it fifty a hundred years later, the reader will get an update on cats and their genes.
What’s inside the cat genome
“Classic” genes of all eukaryotes (higher organisms with nucleu within cellular membrane, which include, apart from cats and humans, all other multi-cellular and numerous unicellular organisms) have similar structure. Their function is the same – to produce a specific protein, which structure is encoded in the gene: each amino acid of the protein under construction corresponds to its own nucleotide triplet.
However, the process of reading and implementing genetic information is very complex. At the beginning of each gene, there are various regulators (on-off switches) of activity, which are switched on and off by special proteins called transcription factors. If the gene is “turned on”, the process of transcription is launched, when a complementary molecule of another nucleic acid, RNA, is assembled on the DNA template. The RNA molecule then “matures” in a process called splicing, when it is split (or sliced) into fragments and sewn together again, turning into mature matrix RNA. It is called so because it serves as a matrix that ribosomes, which are protein “factories”, use as a template to assemble the future protein in a process called translation.
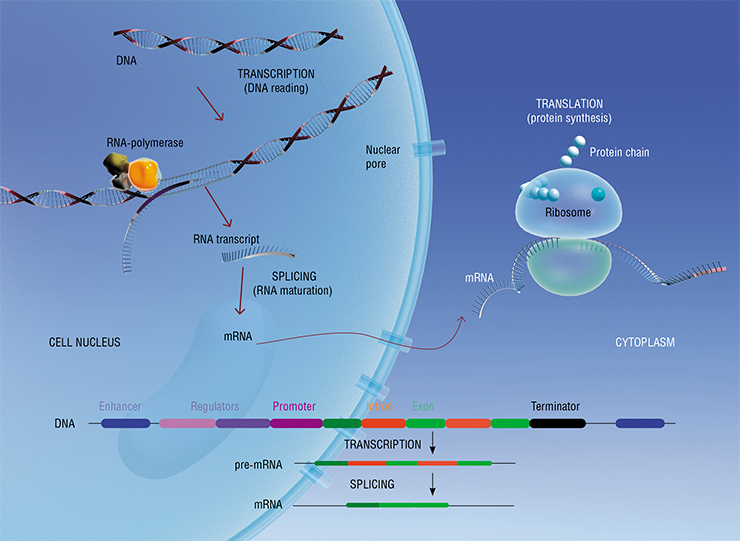
Together the three processes – transcription, splicing, and translation – are regulated by complex interactions of DNA, proteins, and RNA, which depend on the particular cell type, stage of cell cycle, and period of organism development.
For instance, during splicing, some RNA fragments, called exons, are sewn together, and other, called introns, are discarded. The latter normally just degrade afterwards, but sometimes they are used for various purposes, such as regulation of the transcription process itself.
There are mysterious regulatory regions of DNA called enhancers, which play a special role in fine tuning the gene activity. These are very short sequences (50–1500 nucleotides long) that can be adjacent or located very far from the gene whose work they modulate. How enhancers work is still unclear. They are awesome in that they are tissue-specific, i. e. each enhancer only works in a particular part of the body at the particular time.
We now know that the cat genome contains 19587 protein-coding genes. There are also 9438 non-coding genes, which are responsible for the production of non-coding RNA, including transport RNA (tRNA) and ribosomal RNA, as well as a variety of regulatory RNA. Unfortunately, we know virtually nothing about the work of the latter in cats.
In addition, the cat genome contains 494 pseudogenes – non-functioning genomic fossils that were “broken” by mutations. Pseudogenes may look like normal genes, but they are given away by their lack of introns and promoters – the starting points of transcription. This potentially “garbage” group also contains the traces of activity of genomic parasites – retroviruses (a family of RNA-containing viruses, which affect primarily vertebrates). Retroviruses themselves and their fragments, called retroelements, comprise up to one-third of the cat genome. This is a lot, but much less than in the human genome, which is half-retroviral.
The colder the brighter
 There are different kinds of mutations. The most common kind are point mutations, when a single nucleotide is replaced, lost or inserted. As a rule, these mutations occur as errors in the process of replication of DNA, or in the process of reparation, when these and other errors are fixed. About two thirds of these mutations cause changes in the structure of the protein encoded by that gene. Let’s have a look at an example – namely, at albinism, which is complete or partial lack of melanin pigments.
There are different kinds of mutations. The most common kind are point mutations, when a single nucleotide is replaced, lost or inserted. As a rule, these mutations occur as errors in the process of replication of DNA, or in the process of reparation, when these and other errors are fixed. About two thirds of these mutations cause changes in the structure of the protein encoded by that gene. Let’s have a look at an example – namely, at albinism, which is complete or partial lack of melanin pigments.
Coat color in cats, and other mammals, too, is defined by the presence and distribution of two types of pigments in the hairs: black (eumelanin) and yellow (phaeomelanin). Both pigments are synthesized from tyrosine, an amino acid, with the help of an enzyme called tyrosinase. This enzyme is encoded by the gene C (Color). Mutations in this gene either leave the cat without any pigments or disrupt their synthesis in different ways.
There are two point mutations in the C gene (cb and cs), each сaused by a single nucleotide substitution, causing synthesis of tyrosinase with a shift in its temperature optimum. The mutant enzyme doesn’t work well at normal body temperature, but it becomes more effective at slightly lower temperatures. The loss of a single nucleotide in the c and с2 mutations produces the so-called stop codon – a signal terminating transcription. As the result, a truncated, mutant version of mRNA is produced, and a completely faulty, non-functional enzyme is synthesized.
Like humans, cats inherit their genes from two parents, so each gene is present in the genome in at least two copies. If any of the mutations above is inherited only from one parent (such individuals are called heterozygous), the cat will have a normal coat color. One copy of the “correct” gene is enough to supply cells with the necessary amount of melanin. But what will happen if the individual is homozygous – that is, when the cat gets the mutation from both of its parents?
With c or с2 mutations, the cat will be completely white and blue-eyed, because it completely lacks normal tyrosinase. The coat color of cats homozygous for cb mutation is called Burmese, and Siamese – for cs. Siamese cats have strongly colored ears, nose, paws, and tail – i. e. those body parts where the normal temperature is lowered. Burmese cats have some color on the body, albeit paler than normal. We can draw a conclusion that in Burmese cats, the temperature optimum is somewhat higher than in Siamese cats.
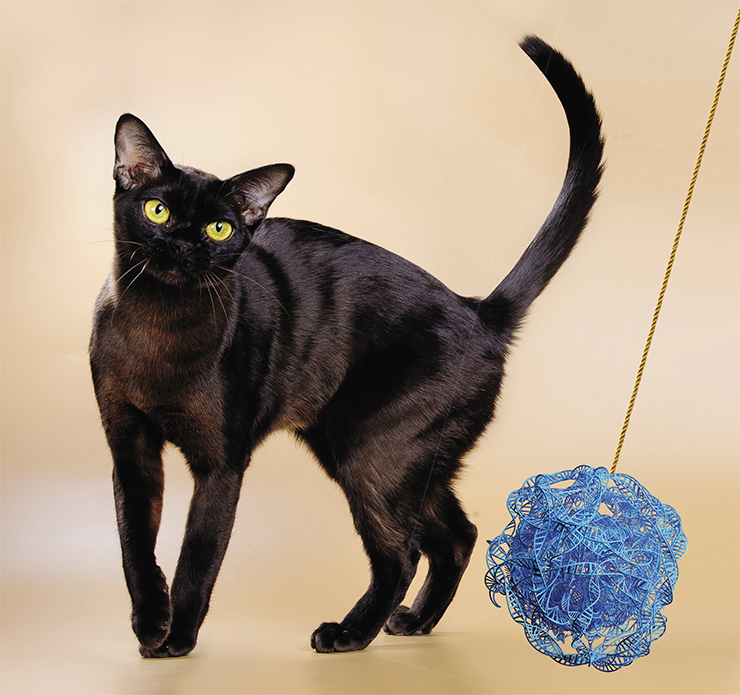

Now here is a question: what color will be a cat with a copy of cs from one parent and a copy of с from the other parent? The correct answer is Burmese. What about a heterozygous cs/cb? This coat color is called Tonkinese – visually, it is something inbetween the Siamese and the Burmese colors. Here, we can infer a simple and clear explanation of the phenomenon of trait dominance. A dominant trait is one which emerges even when a single gene of the corresponding pair is active.
Why are Sphynx cats hairless, and why do Rex cats have curly hair?
Apart from point mutations, cats can have mutations caused by deletion, duplication or insertion of whole fragments of DNA. These mutations often happen when the process of exchange between regions of chromosomes, called crossingover, is disrupted during the formation of gametes.
Several mutations of this kind have been found in cat genes responsible for synthesis and maturation of keratins. These are complex and sturdy fibrillar proteins which make up hair. Disruptions in their structure cause hairlessness (in Sphynx cats) or curly hair (in different types of Rex cats).
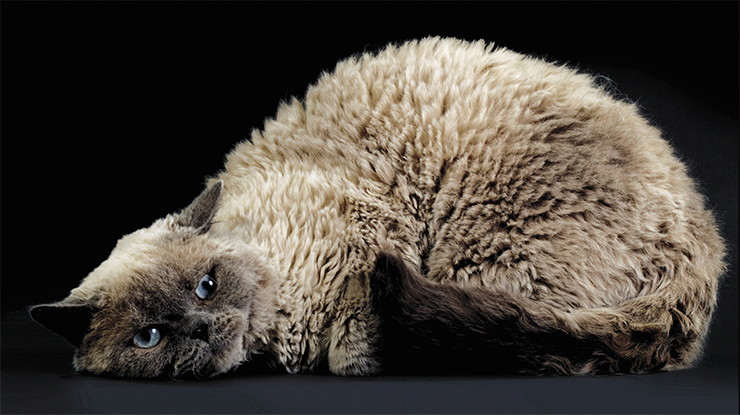
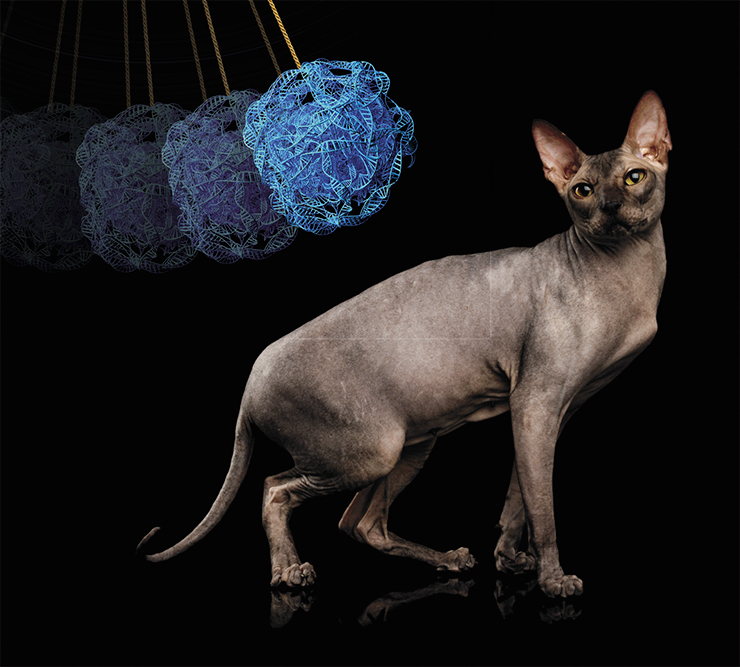


In Devon Rex cats, curly hair is caused by a complex mutation in the KRT71 gene (Gandolfi et al., 2010). The mutation consists in the loss of DNA fragments in two introns and two insertions in one of the exons. As the result of the insertion of extra amino acids, the structure of the keratin changes, compromising its strength. The changes in introns should not be affecting the structure of the encoded protein, because they are snipped out during the maturation of mRNA. However, such mutation can in fact change the activity of the gene by affecting the splicing process and/or stability of the RNA produced from the gene.
The KRT71 gene also has a point mutation – a substitute of guanine to adenosine, creating a stop codon. As the result, a shortened keratin molecule is synthesized, which, through complex interactions with other proteins, leaves homozygous carriers of the mutation – Canadian Sphynx cats – completely hairless. Surprisingly, the complex mutation of the Devon Rex cats “only” curls their hair into ringlets, while a single-“letter” change robs Sphynxes of their fur.
The structure of the keratin can change following mutations in the keratin gene itself as well in genes responsible for the “maturation” of this protein. Selkirk Rex cats owe their curly hair to a mutation in the SADRE gene (Selkirk Autosomal Dominant Rex), which causes a loss of five amino acids in the keratin protein, the product of the KRT71 gene (Gandolfi et al., 2013).
Retrovirus was here
The cat, like many other animals, has mutations induced by retroviral insertion into a gene. They cause white spots on the body.
In general, the size of white spots on the cat body is very variable: from white “gloves” to the whole white cat. It was long thought that the latter phenotype is defined by the dominant white mutation W (White). All other kinds of white spots were thought to be manifestations of semi-dominant mutations in another gene – S (Spotting). However, in 2014 it was discovered that all variants of white spotting are caused by different mutations in the same gene – W (White spotting) (David et al., 2014).

W-encoded protein ensures migration of various embryo cells derived from the neural crest towards their final destinations. These cells include the melanocytes, which produce pigments that give color to hair and irises. Mutations in the W gene disrupt the migration and poliferation of precursors of these cells, but to a varying degree. These mutations are especially fascinating in that they appeared one after another, one from another, as the result of retroviral invasion.
It all began around 3 million years ago, when a retrovirus over 7 thousand nucleotides long wedged in between two introns of the W gene. This caused a partial malfunction of the gene, resulting in the ws mutation. Cells containing the defective protein migrate to their destination slower than they should. The parts of the skin where they fail to arrive on time remain white. In heterozygous individuals, the delay is insignificant, and the spots are small; homozygous cats have bigger white spots.
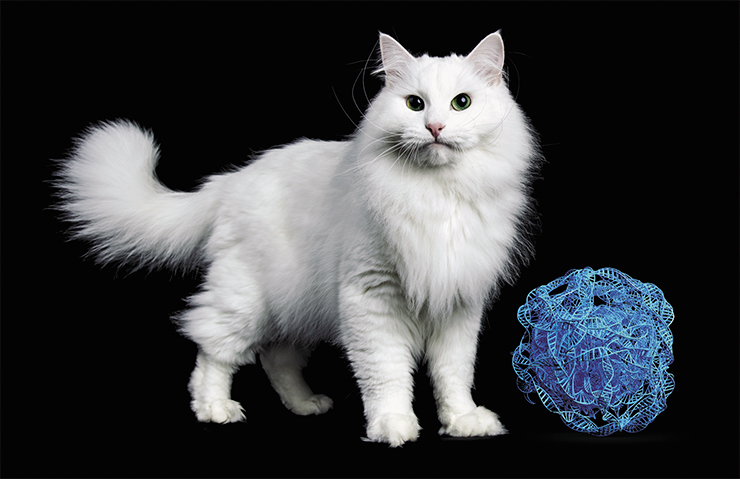

Eventually, the retroelement itself “broke down”, and only a small fragment around 700 nucleotides long remained. This created the dominant white mutation W. However, this did not make life any easier for carriers of the truncated retroelement. Quite the opposite: the new mutation, unlike the previous one, goes beyond slowing the migration and proliferation of the neural crest derivates – it blocks the whole process almost completely.
Many mammals have numerous white spotting mutations. It is remarkable that many of them are caused by mutations in genes homologous to the W gene of the cat. In almost all of these cases molecular analysis showed that these mutations were also caused by retroviral invasion. Each species carried its own retrovirus. What attracts retroviruses to these genes? We do not know.
You need ID? Ears, paws and tails are my ID!
Until now, we’ve been looking at how genes affect coat color and the structure of hair and whiskers. The path from these genes to the corresponding traits is relatively direct and short.
What about genes that control morphological features – paws, ears, tail, or patterns of coat color? Here, the paths are way more complicated, intricate and interesting. They are controlled by a multitude of coordinated genes. The coordination is provided by transcription factors, which are synthesized at the right time and bind to regulatory regions of different genes in the right place, setting off cascades of molecular interactions. Mutations that change the development of these morphological features are usually located in genes coding these transcription factors.
Let’s begin from the tail. Eeyore once complained tearfully that many people misunderstand the tail, lacking in imagination and treating it merely as an additional portion of the back. In fact, these many are technically correct. Vertebrae of the tail and spine derived from the embryonic notochord, and their formation is controlled by the same set of genes.
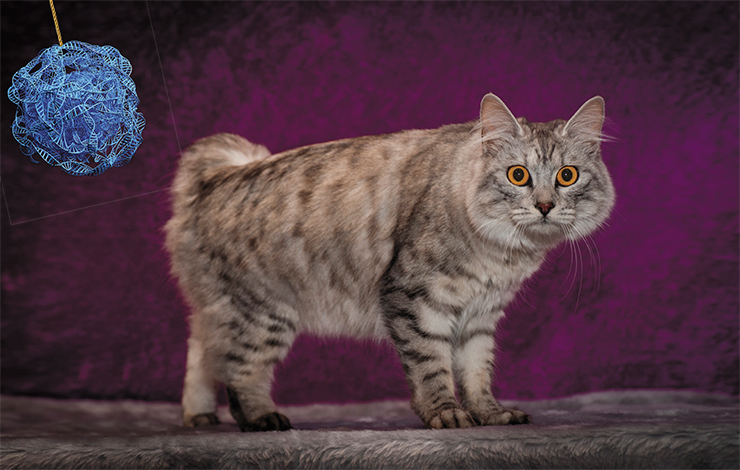

The leading role in these processes is played by a gene called Brachyury. It controls the synthesis of a transcription factor that binds to regulatory regions of many genes involved in the development of the vertebral column, including the neck and tail. The production of this protein begins at the earliest stages of the embryo development, defining the orientation of the head-to-tail and back-to-belly axes. Mutations in this gene have serious consequences.
The tailless cats of the island of Maine and their American relatives have been found to have four mutations in the Brachyury gene: three point substitutes and a duplication combined with a deletion (Buckingham et al., 2013). Each of them leads to synthesis of a shortened protein. Heterozygotes produce enough of “correct” protein to assure normal embryo development, but not the embryonic tail. Homozygotes die on early stages of pregnancy, because the whole function of the gene network dependent on the Brachyury protein goes awry.

The Japanese Bobtail owes its pompom tail to a mutation in the HES 7 gene, which plays an important role in the segmentation of the vertebrate body (Xu et al., 2016). This gene is capable of self-regulation: the encoded protein represses the activity of HES 7 itself. Eventually the protein decays, and its synthesis begins again. This cycle enables the segmentation of the embryo, including the formation of the chain of vertebrae.
The Japanese Bobtail has a point mutation in this gene, which destabilizes the HES 7 protein. The mutant protein decays faster, its cycle gets shorter, and segments turn out very short. Some of them never form completely or even merge together. A normal cat has 22 vertebrae in its tail, while bobtails have 14 to 21 vertebrae. The length of the tail can vary – it is the shortest in homozygotes.
How the Leopard Got His Spots
We think – although we cannot prove – that the same mechanism of activation-inactivation cycle is involved in the formation of repetitive color patterns (stripes, rosettes, and blotches) on the cat body – commonly known as tabby. The genes controlling particular variants of this pattern are well-known. What we do not know is how they work.
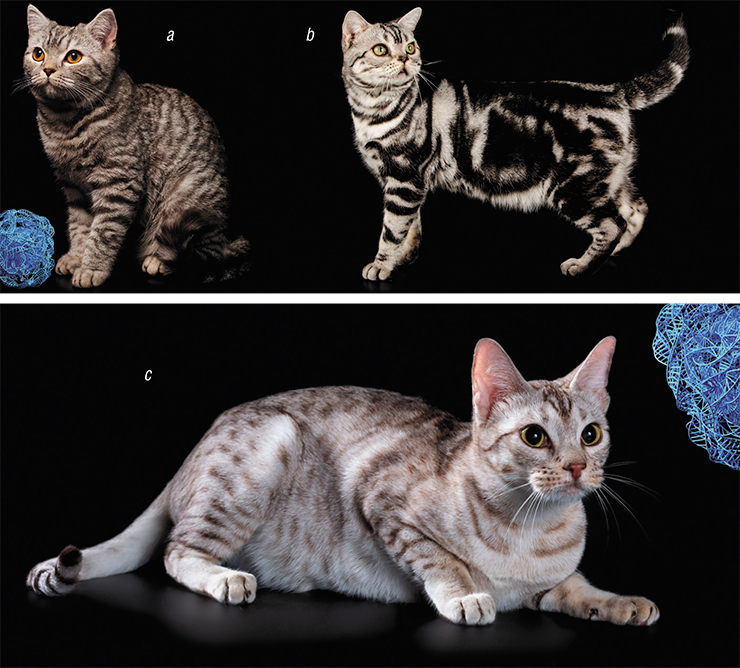

The two most common patterns are mackerel (the usual tiger stripes of the tabby cat) and blotched, with larger elements of the pattern. The mackerel coat pattern is inherited by the domestic cat directly from the ancestors – most modern wild cats have the same pattern. It is controlled by the Ta (Tabby) gene, which encodes an enzyme called transmembrane aminopeptidase Q (Kaelin et al., 2012). This enzyme is involved in controlling diffusion of different substances in intercellular space. There are three known mutations disrupting the structure of this protein, leading to the same result in homozygous individuals – they develop the blotched coat pattern.
We know that in dark-colored areas, black pigment prevails; in light-colored areas, it’s yellow pigment. The proportion of these pigments in a hair depends on the joint action of two gene pairs: the А (Agouti) and E (Extension) pair and the Edn3 and Ednrb pair. We also know that the pattern of the future fur on the embryo body is established 1 or 2 weeks prior to birth. We do not know how it happens. Why do some groups of cells produce dark hair and other grow light fur? How and why do changes in amino acid sequence in the transmembrane aminopeptidase Q cause changes in the relative position of these cells?

It is quite possible that coat pattern formation is affected by the cyclical gene activation-inactivation mechanism. Almost 70 years ago, the great mathematician Alan Turing tried to answer a question asked even earlier by the great poet Rudyard Kipling – how did the leopard get its spots? Turing suggested a simple reaction-diffusion model, where spots and stripes form automatically if the color of each cell depends on the interaction of two diffusing active substances – an activator and an inhibitor. We can (and want to) think that the development of the cat embryo has a point when transmembrane peptidase Q regulates the diffusion of similar substances that regulate pigment production.
Interaction of activators and inhibitors with target tissues guides the development of the cat paws. The order of the future fingers (from thumb or big toe to pinky) depends on the SHH (Sonic Hedgehog) gene, which encodes a transcription factor. The latter participates in the development of limbs as well as lungs, teeth, some structures of the central nervous system, etc. The SHH gene is so important that virtually any mutation in this gene is catastrophic, often resulting in the embryonic death.
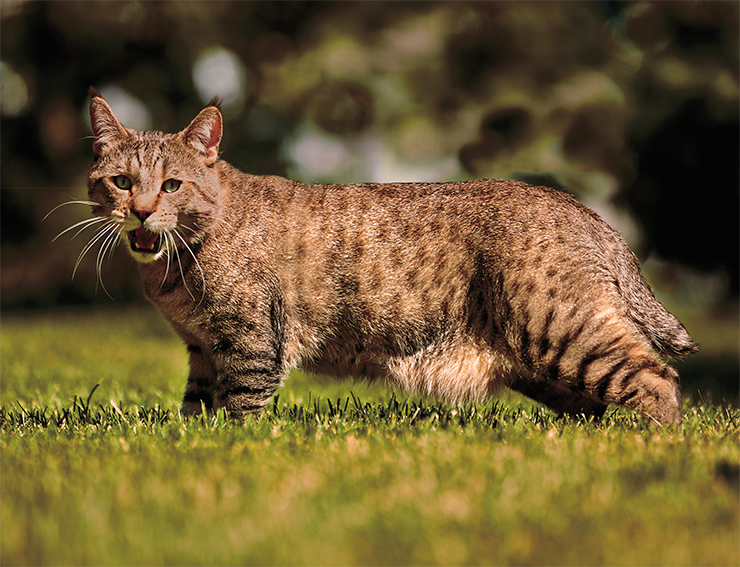

It is apparent that in each of the body structures, SHH works slightly differently, producing the amount of protein required in this particular place. Normally, it is very active around the future pinky finger. Towards the middle of the developing palm (or paw) its activity gradually decreases, and completely stops near the thumb or big toe. The level of SSH activity is controlled by several dozen enhancers. With a mutant enhancer, the gene begins to work actively in some cells of the future thumb, and these cells “decide” they are a separate phalanx and form a sixth finger. This developmental anomaly, known as polydactyly, is fairly common in some cat breeds (Lettice et al., 2008).
Future plans: designer cats
Now that we’ve discussed at the chief achievements of cat genetics of the past decade, let’s try to sneak a peek into the future and guess the key points of “Cats and genes: 50 years later”.
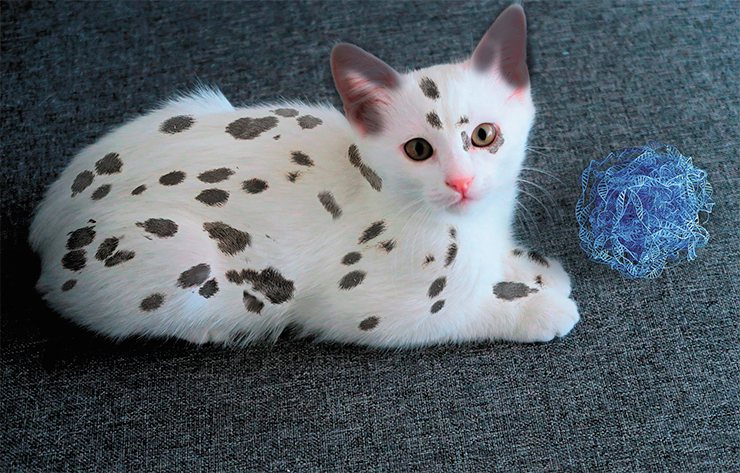

As of today, the International Cat Association (TICA) has accepted 71 cat breeds, the Cat Fanciers Association – 44, the International Feline Association – 43. This does not mean that we can add these numbers up to find out the total number of breeds; all it means that cat lovers can’t agree on that number. Moreover, the question of what constitutes a breed is virtually unsolvable, with so many genetic, breeding and commercial factors and conflicts of interests in play. To put it simply, there are lots of cat breeds. The majority of them were created recently based on separate mutations or combinations of several mutations, with a background of old breed groups.
Today, we are one step away from creating designer cats – animals with predefined qualities. We’ve got all the necessary tools. We know the cat genome, we can clone cats, and we can create transgenic animals. Finally, we’ve got a great tool of gene editing: the CRISPR/Cas9 technology and its modifications allow us to edit the required parts of the genome. The question is, what changes do we want?
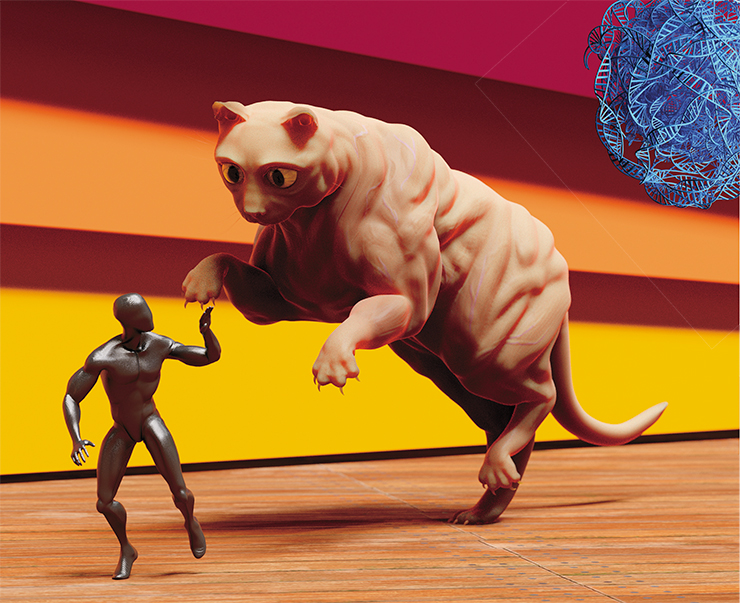

Let’s reverse the question. We definitely do not need changes that will cause suffering to the “gene-edited” animals. We do not want dwarves, “cat pugs”, or giants. Firstly, because they already exist – just look at Maine Coons! Secondly, and more importantly, because body size, like many other measurable traits, are controlled by multiple genes, with each gene’s contribution being very small. Such features are more easily changed by selection. Editing the many genes controlling such traits is an extremely expensive and virtually useless venture.
We must decide what we want and choose our targets. Next, we will make use of Vavilov’s law of homologous variation, stating that similar species have parallel variability, and our knowledge of the genome of the cat and its relatives. Say, the cat does not have the necessary phenotype, but the dog does. What we do is find the dog gene with the necessary mutation, then find the homologous cat gene, and modify it accordingly. Done!
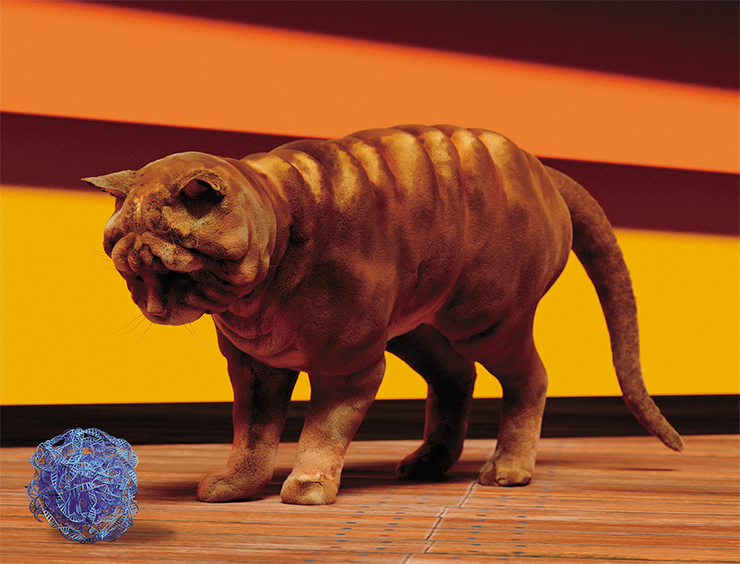
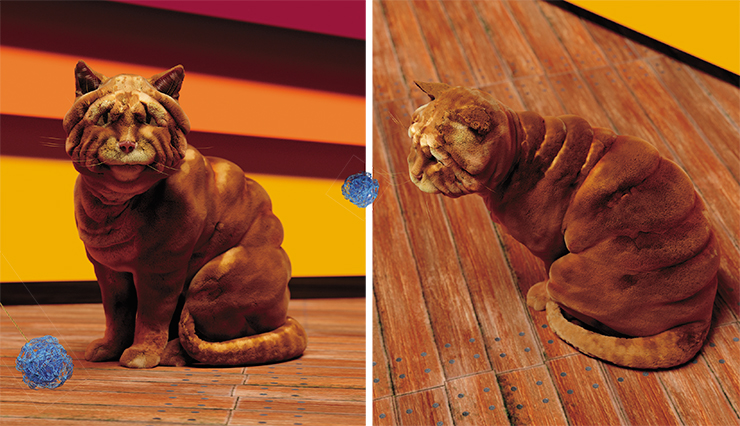


So, we want some previously unseen and unknown cat forms and colors. Shar-Pei cats, Dalmatian cats, bodybuilder cats – like the Belgian Blue bullscattle… Orders accepted! Just kidding. We would never do that ourselves – we’re just outlining the perspectives of the cat genome editing. Let’s get back to the very origins of the unbreakable bond between cats and humans.
The domestication of the cat: where, when, and how
The key events of the Felidae macroevolution were described in detail about 20 years ago. In the past decade, interests have shifted to the more recent feline history: the timeline their domestication and interaction with humans.

Genetic research has shown that all domestic cats have a single ancestor species – the wild cat, Felis silvestris (Driscoll et al., 2009). This species is widely distributed across the Old World: from Scotland to South Africa, and from Spain to Mongolia; it has several subspecies.
DNA-wise, all domestic cats are virtually indistinguishable from the African wildcat, F. s. lybica – a subspecies with a primarily Middle Eastern distribution. African wildcatfossils are known mainly from South Asia and South Europe. Presumably, for millennia, from the Neolithic times to the modern days, these cats have lived primarily in Asia Minor (a part of modern Turkey).
The earliest archaeological evidence of the coexistence of humans and cats was discovered on Cyprus, dating 7—8 thousand years BC. Since no Mediterranean islands, except Sicily, ever had indigenous cat populations, the only way cats could get to Cyprus was together with migrants from the Far East. Apparently, cats were domesticated in the so-called Fertile Crescent about 10 thousand years ago – exactly when the first human settlers came to this legendary Middle East land, often called the cradle of civilization.
Humans created a completely new environment for wild animals. Garbage piles around human settlements provided food and shelter for rodents. The rodents were followed by cats – but not all cats. At first, there was a strict selection of individuals tolerant to human presence, those that could coexist with humans. Why has F. s. lybica become the only subspecies of the wild cat to be domesticated? The answer is the right place, and the right time. As agriculture spread beyond the Fertile Crescent, so did F. s. lybica, keeping the local populations of wild cats from their food resources – i. e. garbage.
Cats conquer the world
Mitochondrial DNA Sequence Analysis of fossils and modern cats allowed to reconstruct the story of their spreading across the world. Modern cats are characterized by five mitotypes – the main variants of the mitochondrial DNA (mDNA) – A, B, C, D, and E, which differ by several mutations (Ottoni et al., 2017).
Almost all (12 out of 14) fossil remains of wild cats dating from 8000 to 800 BCE have the mitotype А. Eventually, cats with this mitotype spread across the whole Old World. It is currently the most common mitotype among domestic cats.
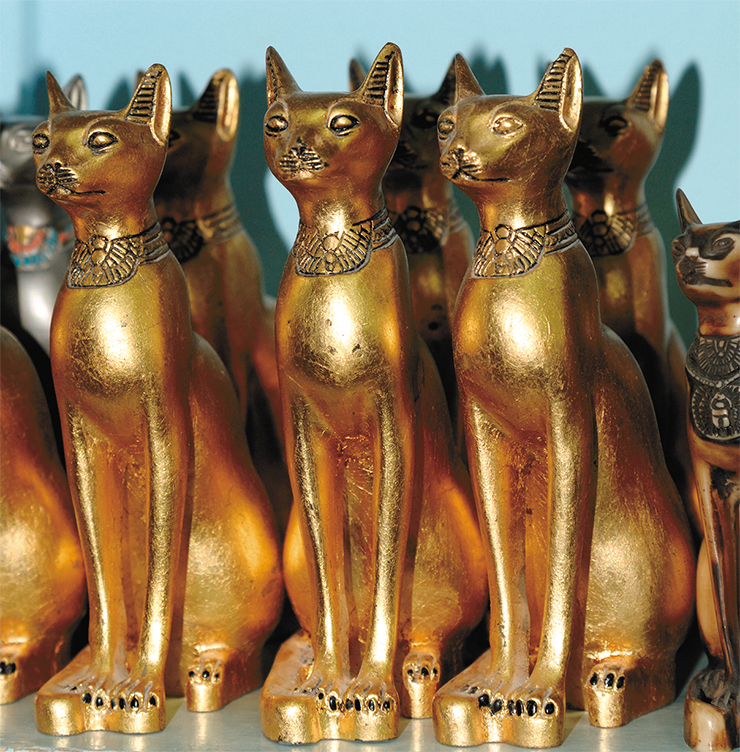
The majority of the discovered mummified Egyptian cats that lived from 7500 to 2000 thousand years ago have the mitotype С. Cats played a major role in the life and religion of the ancient Egyptians. The numerous images and mummies of cats that have reached our days hinted at the possibility that the cat was first domesticated in Egypt. Archaeological and paleontological data do not contradict a theory that the cat was domesticated independently in two locations. However, a hypothesis where the center of domestication was in the Fertile Crescent, with Egypt later becoming a center of cat breeding, seems to be more plausible.
Despite the ban on cat export imposed in Egypt as early as in 1700 BC, Egyptian cats spread across most of the Old World. They were especially popular in Asia Minor: in the first millennium BC, the frequency of the mitotype C in this area was twice as high as that of the mitotype A.
Judging by archaeological and genetic data, the spread of the cat was favored by that of the domestic mouse and the black rat following sea routes, beginning back in the Iron Age. In the Northern Alps, domestic cats appeared shortly after the Roman conquests. During the Middle Ages, cats were a must on ships, which promoted their distribution along military and trading routes. This could explain cat remains dated VII—XI AD with the Egyptian mitotype C in Ralswiek, a Viking seaport (in modern Germany).
People found other uses for cats, other than their main tasks – catching rodents and calming their owners. During the Middle Ages, the cat became one of the telltale attributes of witchcraft, and cat fur was used for garments *.
Moreover, there is still an official national standard #11597–77 “Fur pelts of domestic cat, tanned” that has never been cancelledIn their conquest of new territories, domestic cats interbred with local wild cats, promoting the emergence of two new mitotypes – D and Е. Modern genetic data point at intensive hybridization of feral domestic cats with populations of the European wild cat, putting the latter at risk of extinction.
According to archaeological data, cats came to Siberia with the first Russian migrants. This means that all legends about ancient Siberian cats do not have a scientific basis.

Analysis of geographic distribution of the blotched coat pattern in feral cats allowed reconstructing the migration paths of cats. It is more common in Britain and Iran, and becomes rarer further from these countries. Since the British Empire reigned the seas for a long time, there are still a lot of blotched cats in seaports, especially in the (former) British colonies.
The analysis of ancient nuclear DNA of proto-domestic cats allowed us to check these reconstructions directly. It proved that the blotched coat pattern is relatively recent: the oldest mutant sequence of the Tabby gene was found in cat fossils from South Asia of the Ottoman Empire (XIV—XV AD). Later, its frequency doubled, and it spread across Europe and Africa. In the XVIII century, the blotched pattern became relatively common, and in the following century, it was used to create cat breeds.
Genogeography in the machine learning era
During the last century, “genogeography” was the most thriving branch of cat genetics, by far exceeding all other branches combined by the sheer number of publications. Enthusiasts traveled the world, counting cats of different colors, drew maps, calculated indices and built beautiful models. In the new age of genetics, this profession has come to a halt. A hundred cats were counted in each major city of the world, thus skimming the most interesting material, with only a few minor topics remaining to be covered. Does this mean genogeography is dead? Absolutely no. We see two major paths it could follow next.
As modern genotyping methods become cheaper, molecular genogeography projects develop – and get the necessary funding. Comparing genetic diversity of stray cats from different parts of the world on the one hand, and of the indigenous and immigrant human populations, on the other hand, will give us insight on the evolution of the relationship between the two unique species. Darwin could have written, prophetically, that “light will be shed upon the migration of man and cat and their shared history”.
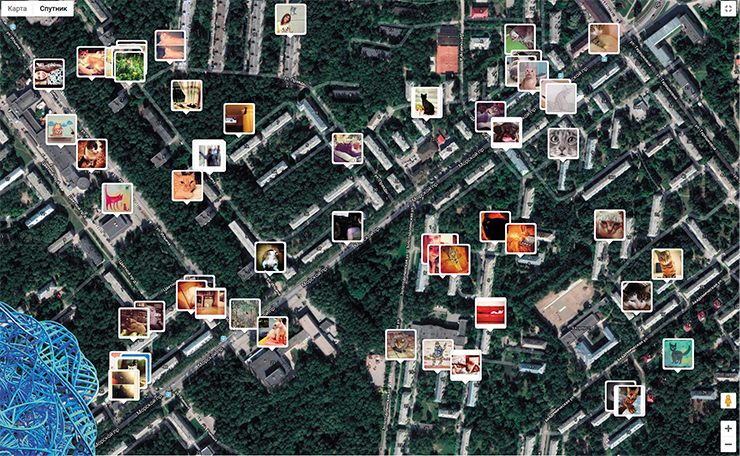

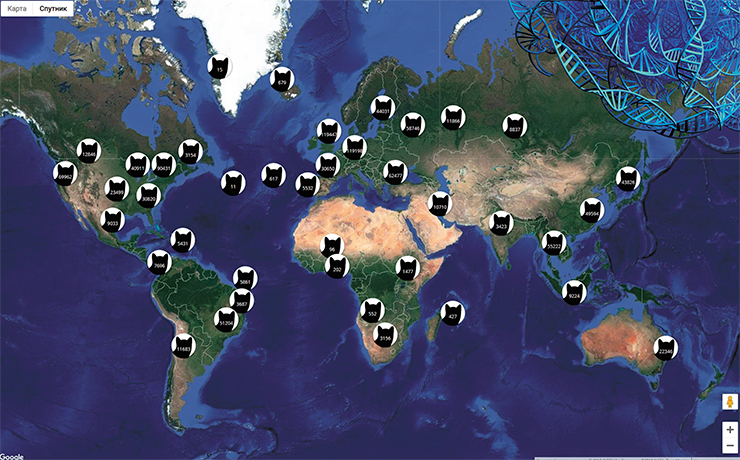
The second path assumes the use of social networks and citizen science to solve genogeographical problems. What is the most posted subject in the world? That’s right, cat pics! All social networks are flooded with cats. Are there ways to put this treasure trove to good use? There are! We need an API that would fish out these cats from the Net. Such an interface has already been written – it is building a global map of cat pics, available at https://iknowwhereyourcatlives.com/. There, you can see locations and photos of all cats with geotagged photos.
The next step here is the creation of a neural network that would define the phenotype of the posted cats. This task is more challenging than it seems. GooglePhoto can recognize human faces in front and side views and a broad age range, and can tell cats from humans, and it can surely be taught to recognize cat faces: this one here is Vas’ka, and that one is Jean-Jacques. However, GooglePhoto is bad with colors and textures. Our programmer friends are struggling with the task of teaching the neural network to tell blue cats from red cats. Perhaps, we just need different programmer friends? In any case, we are absolutely certain that solving this task is a matter of time, and we will get next-generation genogeographic maps.
Finishing his “Origin of Species”, Darwin wrote: “… we can dimly foresee that there will be a considerable revolution in natural history”. Finishing this article, we dimly foresee the enormous progress in all areas of genetics, which will most certainly include the genetics of the cat.

New amazing breeds of cats will be created. New methods of genetic and genomic editing will appear. The mystery of the blotched pattern will be solved. We will understand how and to what extent do genes control development, and how they interact with one another, making possible the development of our cats – so different, so amazing, and so wonderful.
People will learn to reconstruct genomes – and the appearance, and even behavioral features – of extinct animals. We will learn if the saber-toothed cat could purr, and if it liked valerian drops.
There will be a lot to write about in “Cats and genes: fifty years later”.
References
Buckingham K. J., McMillin M. J., Brassil M. M. et al. Multiple Mutant T Alleles Cause Haploinsufficiency of Brachyury and Short Tails in Manx Cats // Mammalian Genome. 2013. V. 24. P. 400–408.
David V. A., Marilyn Menotti-raymond, Wallace A. C. et al. Endogenous Retrovirus Insertion in the KIT Oncogene Determines White and White Spotting in Domestic Cats // G3-Genes Genomes Genetics. 2014. V. 4. P. 1881–1891.
Driscoll C. A., Juliet Clutton-Brock, Kitchener A. C. et al. The Taming of the Cat // Scientific American. 2009. V. 300. P. 68–75.
Gandolfi B., Alhaddad H., Joslin S. E. K. et al. A Splice Variant in KRT71 Is Associated with Curly Coat Phenotype of Selkirk Rex Cats // Scientific Reports. 2013. V. 3. P. 2000.
Gandolfi B., Outerbridge C. A., Beresford L. G. et al. The Naked Truth: Sphynx and Devon Rex Cat Breed Mutations in KRT71 // Mammalian Genome. 2010. V. 21. P. 509–515.
Kaelin C. B., Xu X., Hong L. Z. et al. Specifying and Sustaining Pigmentation Patterns in Domestic and Wild Cats // Science. 2012. V. 337. P. 1536–1541.
Lettice L. A., Hill A. E., Devenney P. S. et al. Point Mutations in a Distant Sonic Hedgehog Cis-Regulator Generate a Variable Regulatory Output Responsible for Preaxial Polydactyly // Human Molecular Genetics. 2008. V. 17. N. 1. P. 978–985.
Mosher D. S., Quignon P., Bustamante C. D. et al. A Mutation in the Myostatin Gene Increases Muscle Mass and Enhances Racing Performance in Heterozygote Dogs // PLOS Genet. 2007. V. 3. P. 1–8.
Olsson M., Meadows J. R. S., Truvé K. et al. A Novel Unstable Duplication Upstream of HAS2 Predisposes to a Breed-Defining Skin Phenotype and a Periodic Fever Syndrome in Chinese Shar-Pei Dogs // PLoS Genetics. 2011. V. 7. N. 3.
Ottoni C., Neer W. V., Geigl E. et al. The Palaeogenetics of Cat Dispersal in the Ancient World // Nature Ecology & Evolution. 2017. V. 1. N. 139.
Xu X., Sun X., Hu X. et al. Whole Genome Sequencing Identifies a Missense Mutation in HES7 Associated with Short Tails in Asian Domestic Cats // Scientific Reports. 2016. V. 6. N. 31583.