Cats and Genes: 30 Years Later
What has happened in the genetics of the domestic cat over the last 20 years? The first results on the cat genome sequencing appeared in 2007. So far about 65 percent of the cat genes have been sequenced. Comparison of the cat genome with the well studied human, chimpanzee, dog, cow, mouse, and rat genomes allowed 20 285 cat genes to be identified. Consequently, the total number of cat genes is likely to be close to the number of human genes, i. e., about 30 000.
Comparison of the genetic content of the cat chromosomes and chromosomes of other mammals gave very interesting and quite unexpected results. In particular, cats, as well as humans, have rather insignificantly rearranged its chromosomes over the 80-90 million years of evolution from a common ancestor.
The beginning of this story dates back to my student time at Novosibirsk State University. Once our English teacher asked us to retell a scientific paper that would be interesting to the other students in the group. I was looking through journals, and the cover photo of the Journal of Heredity with a fountain in a city square caught my attention. There, around the fountain, 94 cats were sitting, lying, and walking! The photo referred to the paper titled “Some Cats from Sao Paulo, Brazil”, one of a series of papers on the gene geography of cats.
Cats are an ideal object for such studies, because their populations contain a high frequency of mutants for various genes determining coat color (gray, black, white, red, spotted, and so on). As long ago as the late 1940s, J.B.S. Haldane, an outstanding British geneticist, paid attention to this phenomenon and initiated geneticists worldwide to count cats. This founded the outlines of a worldwide map of cat gene geography. Comparison of different populations with respect to the frequencies of coat color genes elucidated the evolution of cats and factors that had determined it, namely, natural and artificial selection, migration, isolation, and genetic drift.
This map had one disadvantage – our entire country (at that time, the USSR) was a terra incognita. Naturally, I commenced filling this gap. First, I counted cats in Akademgorodok and then, in all the cities of our vast country where my life took me.
Cat counting itself was very exciting, adventurous, and even somewhat risky. I had to take into account not only animal behavior, but also public attitude, as a person spying and ferreting out in the yards and breezeways caused a just suspicion. At best, the locals took this person for a tax officer and at worst, for a foreign agent. Especially, when this person every now and then took out a notebook and started to put something down, the strengthening suspicions could cause most adverse consequences.
Therefore, I had to observe and record cats without getting attention, preferably, in motion without slowing down. When my memory “accumulated” more than five cats, I used to enter a pay phone booth, take off the receiver, and ask an imaginary Mariya Ivanovna to tell the phone number of a similarly imaginary Ivan Ivanovich. Than I calmly put down the accumulated information, thanked Mariya Ivanovna, and rang off.
I then tried to expand the scale of my gene geography research by recruiting multimillion cohorts of Soviet schoolchildren. For this purpose, the USSR Central Television shot an educational movie, in which the most beautiful TV anchor Irina Ermilova and I walked around Moscow pretending to come accidentally upon cats. I diagnosed them according to their genotypes, and Irina put down the data.
In fact, the project was just a contrived coincidence. The director mobilized teachers of biology from several Moscow schools. In turn, they mobilized their pupils, gave each of them a vial of valerian drops, and sent them to hunt for stray cats. A TV van brought Irina and me to a site. Here we assumed thoughtful poses (she, all in red, and I, all in white), the director uttered “Action!”, and another schoolchild released another cat towards us.
The map of marbled coloration distribution in cat populations resembles a diffusion pattern. The concentration is maximum in the British Isles, slightly lower in France, and then gradually decreases as the distance from the center of distribution becomes longer.A frequency of marbled cats in seaports, in particular, Vladivostok and Petropavlovsk-Kamchatski, indicates a sea-going route of their distrtibution. In Siberia marbled cats are exclusively rare
After this, schoolchildren from all over the country sent me three large bags of letters with descriptions of local cat populations. These letters were admirable but poorly informative from a scientific standpoint. Thus, I had to recruit experts, my colleague geneticists.
The fruit of this work was two scientific papers in the mentioned Journal of Heredity and the monograph Cat Genetics. Being fascinated by this issue, I wrote two popular science books, Essays on Mutants (1983) and Cats and Genes (1995).
In addition, I wrote two popular science papers for the journal Khimiya i Zhizn’ (Chemistry and Life): the first paper titled “Cats, Genes, and Geography” appeared in 1979 and the second, “Cats and Genes: Ten Years Later”, in 1989. Thus, this paper had to be titled “Cats and Genes: 30 Years Later”.
Cat chromosomes
What has happened in the genetics of the domestic cat over the last 20 years? On the one hand, the situation with gene geography practically has not changed. The global maps became informative enough as early as 20 years ago, and new data did not add to their interpretation. Consequently, publications on this issue eventually faded away. On the other hand, important events have occurred in the cat genetics over the last decade, in particular, those connected with the development of total genome sequencing methods.
The first results on the cat genome sequencing appeared in 2007. So far about 65 percent of the cat genes have been sequenced. Comparison of the cat genome with the well studied human, chimpanzee, dog, cow, mouse, and rat genomes allowed 20 285 cat genes to be identified. Consequently, the total number of cat genes is likely to be close to the number of human genes, i. e., about 30 000.
Comparison of the genetic content of the cat chromosomes and chromosomes of other mammals gave very interesting and quite unexpected results. In particular, cats, as well as humans, have rather insignificantly rearranged its chromosomes over the 80—90 million years of evolution from a common ancestor.
However, although the cat chromosomes look extremely conserved in a macroevolutionary context, they appeared to be the champions among the studied mammalian species in the frequency of genetic recombination, the redistribution of genes through the exchange of chromosomal regions between chromosome pairs. As is known, recombination is the main supplier of new gene combinations, which feed for natural selection as well as the micro- and macroevolutionary processes.
GENETIC RECOMBINATION AS A SOURCE OF HEREDITARY VARIATION
Recombination takes place in the first division of meiosis, the process comprising two divisions of a cell and leading to formation of four germ cells containing a single rather than double chromosome set.When preparing for the first division, homologous (paired) chromosomes are juxtaposed and aligned, and multiple double-strand breaks are formed in their DNA. The protein Rad51 is an active player in the repair of DNA breaks: it binds to the free ends of broken DNA strands and inserts them into the DNA of a homologous chromosome concurrently unwinding the DNA target.
Having found the complementary region, the inserted DNA strand pairs with it. However, the majority of links between DNA fragments are cleaved and linked to restore the initial state of the DNA strands (exchangeless pathway). In all the studied mammals (except for the cat!), less than ten percent of the breaks are linked in a crossover manner (exchange pathway). In this process, the DNA strand of one homologous chromosome is linked at exchange points to the DNA of the other homolog. These are the recombination points.
The marker for recombination points is the protein MLH1, a member of the repair protein family whose function is to repair DNA mispairing, i. e., to excise the unpaired nucleotides. The antibodies to MLH1 labeled with fluorescent dyes provide markers for analyzing the frequency of recombination events and their distribution over the genome
As has been found, cat heads the list of all the studied mammalian species in the density of recombination events, i. e., exchanges of individual fragments between DNA strands per unit chromosome length. The mean distance between the recombination points in cat chromosomes is 3.7 microns (versus 7.1 microns in mice and 6.0 microns in humans). In addition, the lower threshold for this distance is as small as 0.05 micron, which is close to the resolution of a microscope.
A high recombination density in the cat chromosomes results from a high recombination efficiency. In all the studied mammals, only a small part (less than 10 percent) of the primary bonds between the DNA of homologous chromosomes is linked in a crossover manner, thereby forming recombinant chromosomes. As for cats, the rate of the recombinationally resolved cross-links reaches 25 percent. This shows that the recombination process in cats is organized in a more economical manner as compared with the other mammals: it provides for a sufficiently high recombination level on the background of a smaller number of double-strand breaks.
Siamese substitution
Two decades ago, the cat genetic map contained only several dozens of genes, whereas their current number reaches two thousand. In particular, the genes determining coat color have been sequenced and mapped, and critical points mutations in which change coloration have been determined.
In particular, two coloration mutations are localized onto one somatic (non-sex) chromosome. Mutation of the dominant white color, located within the c-kit protooncogen, interferes with the migratory ability of melanoblasts, precursors of embryonic pigment cells. Since melanoblasts fail to reach the skin in time, the hair remains without the pigment. Consequently, the fur grows completely white. However, if sometimes melanoblasts succeed in entering the hair follicles on the cat’s head, small colored patches develop there. In the carriers of this mutation, the number of melanoblasts that have reached the retina can be different. If this number is large, the eyes will have a standard yellow color; otherwise, the eyes will be blue.
The same chromosome carries the gene determining the coloration pattern. The normal allele (structural form) gives a tabby, tiger-like coloration. Sometimes these strips are continuous and sometimes, dashed. We know the semidominant mutation Abyssinian tabby. The homozygotes (i. e., individuals carrying a pair of the same alleles) for this mutation display no stripes on their coat: the animals have a uniform coloration. As for the heterozygotes for this mutation, they have strips on the tail, muzzle, and legs. The recessive mutation in the same gene, marble tabby, transforms the transverse strips into curly or irregular-shaped spots. Usually, such cats have a wide black band long the backbone.
In the case of albinism, a widespread phenomenon in various mammalian species, mutations take place in the gene encoding the enzyme tyrosinase. These mutations either completely block the synthesis of this enzyme or result in a defective enzyme with an altered activity.
Several mutations of this kind have been described in cats. Tyrosinase activity in the homozygotes for the mutation Burmese albinism is somewhat decreased as compared with the norm. Moreover, the degree of enzyme activity inhibition depends on body temperature: the enzyme is more active at a temperature below the normal level. This is why the Burmese cats display a more intensive fur coloration on paws, nose, and points of the tail and ears, i. e., on the body parts with a decreased temperature,
The same is true for the mutation referred to as Siamese albinism. However, its carriers display a higher degree of depigmentation: as a rule, the body fur of Siamese cats has no pigment, whereas paws, nose, and points of the tail and ears retain color. However, even these parts are less pigmented as compared with Burmese cats. The eyes, as a rule, are blue, because of a decreased pigment content in the retina.
Today we know precisely the molecular nature of these mutations: they result from the substitution of only one nucleotide in the sequence of the corresponding gene! In Siamese cats, the nucleotide at position 422 from the beginning of the gene coding for tyrosinase is changed. In contrast to common cats having guanine at this position, Siamese cats have adenine. Consequently, the nucleotide sequence encoding the amino acid arginine is transformed into the sequence determining another amino acid, glycine. The substitution of glycine for arginine in the tyrosinase protein leads to a decrease in the corresponding enzymatic activity at a normal body temperature.
In the Burmese cats, an analogous fading of the coat color on the body is determined by a nucleotide substitution at position 227.
A relative of horse
A detailed analysis of the genomes of cats and other mammals suggested a radical revision of the overall mammalian phylogenetic tree. The previous phylogenetic tree looked like a bush rather than a tree, with all the branches representing orders coming from one root. The modern phylogenetic tree displays successive branchings.
The three main branches are Afrotheria (elephants, sea cows, hyraxes, aardvarks, golden moles, etc.), Edentata (endemics from South America – armadillos, sloths, and anteaters), and Laurasiatheria (all the remaining eutherian mammals). These three stems formed due to splitting of the ancient supercontinent Pangaea into Gondwana to form Africa, India, South America, Antarctica, and Australia and Laurasia to form Eurasia and North America. Then Gondwana separated into the corresponding continents, and Africa was the first to break away. It was on this isolated continent that the superorder Afrotheria developed.
Cats, the focus of our interest, belong to the order of carnivores, located in the branch of Laurasiatheria, which contains the largest number of species. Among other things, further branching led to the formation of the group uniting carnivores, ungulates, and chiropters (bats). Bizarre as this group may look, its common origin has been convincingly confirmed by molecular data. Moreover, these data demonstrate that further branching within this group followed a pattern different from the one that could be inferred from the external appearance of the animals the group comprises.
The group of Cetartiodactyla was the first to diverge. (This is not a misprint: this group unites the whales (Cetacea) and even-toed ungulates (Artiodactyla). According to the old “premolecular” phylogenetic tree, whales originated directly from the root of the mammalian phylogenetic “bush”. As for the current views, the closest relative of whales is hippopotamus.) The other branch, Pegasoferae, divided to form odd-toed ungulates (horses, tapirs, and rhinoceroses), carnivores (cats, dogs, bears, walruses, and so on), and chiropters (bats).
The branching order within the superorder Pegasoferae is still unclear; however, certain data suggest that chiropters were the first to separate followed by the divergence of odd-toed ungulates and carnivores. What is known for sure is that the last common ancestor of horses and cats existed later (i. e., closer to our time) than the last common ancestor of horses and cows.
The feline phylogenetic tree has also been revised considerably over the last two decades. It has been found that the first branching in this family took place about 11 million years ago (MYA) in Asia, with separation of the lineage of large roaring cats (lions, tigers, leopards, jaguars, and snow leopards). Many species belonging to this group have practically identical chromosome sets. In nature they remain as separate species; however, in captivity they readily produce hybrid offspring. Many zoos have tiglions, ligers, and other hybrids. Although the majority of these hybrids are sterile, the mere possibility to obtain viable hybrids between these species suggests a close genetic similarity between the large roaring cats.
The second group, which also separated in Asia, comprises marbled cats and Asian golden cats, now inhabiting Southeast Asia. The branch that separated from this lineage and migrated to Africa contains servals, caracals, and African golden cats. This occurred 6—10 MYA, when the world ocean was rather shallow, while Africa and Asia were connected by an isthmus in the region of the modern Red Sea.
WHY CARBON COPY HAS NO ORANGE PATCHES
As is known, she-cats, similarly to other mammalian females, have two X chromosomes in their genome. This is where the gene determining the coat coloration is located. This gene has two alleles, which determine orange and not orange (black) color.Both X chromosomes are active in zygote, the cell produced by the fusion of egg and spermatozoid. However, in the course of cell divisions and subsequent differentiation, one of the X chromosomes is inactivated in all the body’s cells, including the future pigment cells. If a cat is heterozygous for the coloration gene, the chromosome carrying the allele conferring orange color can be inactivated in part of the cells and the allele conferring black color, in the others. The daughter cells stringently inherit the state of the X chromosome. Thus, a tortoiseshell coloration is formed.
Presumably, during cloning the inactivated X chromosome failed to reactivate completely in the nucleus of the reconstructed egg cell extracted from a common somatic cell.
During the same time, the remaining cats colonized Asia, and part of them entered North America via the Bering Bridge. This is the region where the most ancient puma, ocelot, and lynx fossils are found. Later, descendants of the North American cats migrated back to Asia and then to Africa to give birth to Eurasian lynx and African cheetah. In the late Pliocene (2—3 MYA), the Isthmus of Panama was formed between North and South Americas. The ocelot lineage entered South America to give rise to seven new feline species. Puma and jaguar also migrated there from North America. The remaining Asian cats partitioned into individual genera and species in Eurasia during the last 5 million years. This is the group that contains domestic cats.
How many lives does a cat have?
As is known, the first cloned animal was a sheep: the famous Dolly was born in 1996. Five years ago, the first cloned cat appeared, which was wittily named CC (Carbon Copy).
A tortoiseshell (gray-orange) cat with a white patch, named Rainbow, was chosen as the template for cloning. Both egg and somatic cells were isolated from its ovaries. The nucleus was removed from each egg cell and replaced with the nucleus isolated from a somatic cell. After electric stimulation, the reconstructed egg cells were transplanted into the uterus of a gray tabby cat. This surrogate mother gave birth to CC (Shin et al., 2002).
In its genotype, CC was a precise copy of Rainbow; however, CC differed from it in the external appearance, because it lacked orange patches. The authors of the mentioned paper in the Nature explained this in a rather equivocate manner, stating that the pigmentation pattern in multicolored animals resulted not only from genetic factors, but also from developmental factors not controlled by the genotype.
This statement can be understood in many different ways. This is how I understand it. In the somatic cell that donated the nucleus used for creating CC, one of the two sex X chromosomes was inactivated, actually, the one that carried the allele determining a orange coloration. As is known, the state of the X chromosome is stably inherited in the generations of somatic cells.
What is amazing about CC is that the transfer of the nucleus from a somatic cell to an egg cell failed to induce a reactivation of the X chromosome. Consequently, the cloning procedure did not lead to a complete reprogramming of the nucleus. Presumably, this is the cause of health and reproductive problems in cloned animals. However, CC has no heath problems. It is now eight years old, and three years ago it gave birth to three kittens.
The company Genetic Savings & Clone, which funded creation of CC, tried to do business out of it. However, the things went wrong, as they succeeded in selling only two cloned pets (for US $50 000 and $32 000), and that was that.
Nonetheless, the baton was picked up by South Korea: the first cat was cloned there in 2004. Korean researchers regarded this cloning not as an end in itself but rather as an intermediate stage in solving another, much more ambitious problem.
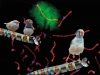
They were interested in producing GM (genetically modified) cats. For this purpose, they isolated fibroblasts, cells of the connective tissue from the ear of a white angora cat. The fibroblasts were cultivated in the nutrient medium containing a mobile genetic element carrying the gene coding for red fluorescent protein.
After this mobile element entered the fibroblast nuclei and inserted into the host DNA, these nuclei were isolated and transferred into enucleated eggs. Two GM kittens developed from these reconstructed egg cells; the red fluorescent protein was synthesized practically in all the cells of their body. Consequently, these cats radiated mysterious red light when exposed to ultraviolet radiation (Yin et al., 2008). The most intriguing is the last phrase in the paper published in the Biology of Reproduction to the effect that this technology will be useful for a directed creation of design cats.
It is difficult to imagine what will be the price for such design GM cats and who will need them. As for me, I am in no way afraid of genetically modified cats. Indeed, giving it a second thought, all the cats in the world have been genetically modified over the endless generations of natural and artificial selections of their ancestors.
P. S. Remembering well the consequences of my science popularization activities, I would like to inform that I do not clone or genetically modify cats, give no recommendations on cat mating, and will not take any kittens.
References
Borodin P. M. Essays on Mutants. – Moscow: Znanie, 1983.
Borodin P. M., Ruvinsky A. O. Cat Genetics. – Novosibirsk: Nauka, 1992.
Borodin P. M. Cats and Genes. – ZAO Zoosalon, 1995.
Borodin P. M. Cats, Genes, and Geography. // Khimiya i Zhizn’ (Chemistry and Life). – 1979. No. 4. P. 40—46.
Borodin P. M. Cats and Genes: 10 Years Later. // Khimiya i Zhizn’ (Chemistry and Life). – 1989. No. 4. P. 40—45.
Shin T, Kraemer D, Pryor J, Liu L, Rugila J, Howe L et al. A cat cloned by nuclear transplantation. // Nature. – 2002. V. 415(6874). – P. 859
The work on analysis of recombination in cats was supported by the Russian Foundation for Basic Research (grant no. 04-04-48024-a)
The author and editorial board are grateful to I. Bodunov (Cat Fancier’s Club Asia, Novosibirsk, Russia), M. Westhusin, and L. Wadsworth (Texas A&M College of Veterinary Medicine, College-Station, United States) for their assistance in preparing illustrations