Padlocked DNA
DNA is one of the treasures of a living cell, which provides for its regular function. The emerging lesions in DNA molecules are “fixed” by specialized repair systems involving ensembles of protein enzymes. Despite such efforts, at any time any cell contains tens of thousands of manifold DNA lesions, one of which may be enough for cell death.
However, in the case of malignant tumors, successful performance of the repair system correcting DNA damage due to cancer therapy may bring to naught the treatment efforts. Siberian researchers have developed a new method for detecting the Ku antigen, a key protein of the system that repairs the DNA double-strand breaks caused by tumor radiotherapy. That is why such studies of DNA repair mechanisms are highly important for practical use as well as basic research
DNA is one of the treasures of a living cell, which provides for its regular functioning. The constantly emerging lesions in DNA molecules are “fixed” by specialized repair systems that involve ensembles of protein enzymes. Despite such efforts, any cell at any moment contains tens of thousands of manifold DNA lesions. Although a single lesion may be sufficient for cell death, the estimate of the situation becomes opposite in the case of a malignant tumor cell. In this case, successful performance of the repair system to correct the DNA damage may bring to naught the treatment efforts. That is why the insight into DNA repair mechanisms, including knowledge about the proteins involved in repair, has both theoretical and practical value
DNA—the major genetic information repository of living beings—is constantly exposed to various factors of chemical and physical nature, such as free radicals and ultraviolet radiation. This causes DNA damage, including the loss and oxidation of nitrogenous bases, cross-linking between DNA strands, and even breaks in the strands.
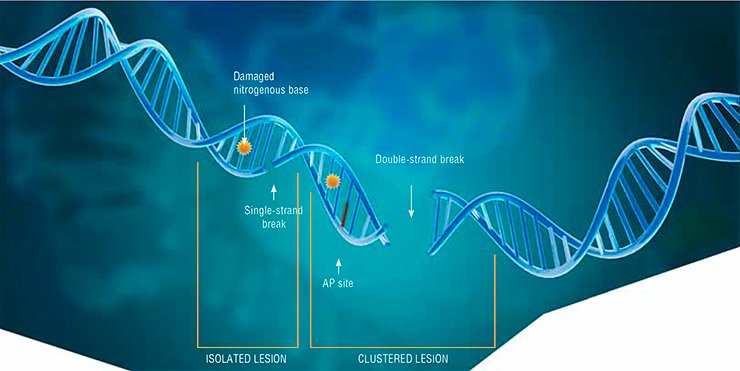
The most dangerous type of DNA damage is double-strand breaks. Since DNA is a helix of two complementary nucleotide strands, breaks in one of the strands, as a rule, do not cause any catastrophic consequences for the cell: the second strand carries all the necessary information for the complete repair of the lesion. However, even a single double-strand break may lead to cell death, while the incorrect repair of such a lesion may result in mutation or loss of genetic information. If several such lesions emerge concurrently and at a short distance fr om one another, they actually make DNA fall into pieces.
Other types of DNA damage may also be dangerous for cell life if they are located in close vicinity, because the repair process necessarily includes a stage when the DNA strand is broken.
Such grouped (clustered) lesions are characteristic of ionizing radiation, which is why it is used to destroy cancer cells. The types of radiotherapy differ in the degree of the DNA structure damage they cause and in the patterns of induced lesions, tumor cells being the most sensitive to ionizing radiation since they divide more actively as compared with healthy cells.
However, this anticancer therapy is not always effective, which can be explained by the high activity of the cell proteins that repair double-strand breaks. In this perspective, the methods allowing for a decrease in the activity of “repair machines” and thus for annihilation of malignant cells are of great practical interest.
Evidently, efficient control of the repair (DNA “fixing”) systems requires precise information about all the proteins involved in DNA repair, including their quantitative patterns in the cell and their part-time duties, i.e. their roles in other cellular processes. This is extremely important since the body, as a whole, may be irreparably injured if the cell systems that provide for DNA repair are suppressed.
All this is fully applicable to our “hero” Ku antigen, the protein with many talents and a hard life.
Repairing DNA
To understand the functions and role of Ku antigen, first it is necessary to get acquainted with two major mechanisms underlying the repair of DNA double-strand breaks, namely homologous recombination and nonhomologous end joining.
The first mechanism is mainly used to repair double-strand breaks that emerge in an actively dividing cell at the phases of cell cycles when copies of the DNA molecule have already been synthesized. These copies are used as a template for repeated DNA synthesis at the site of damage. During this process, implemented by a specialized protein ensemble, no genetic information is lost or distorted thanks to the available undamaged DNA molecules identical to the initial ones.
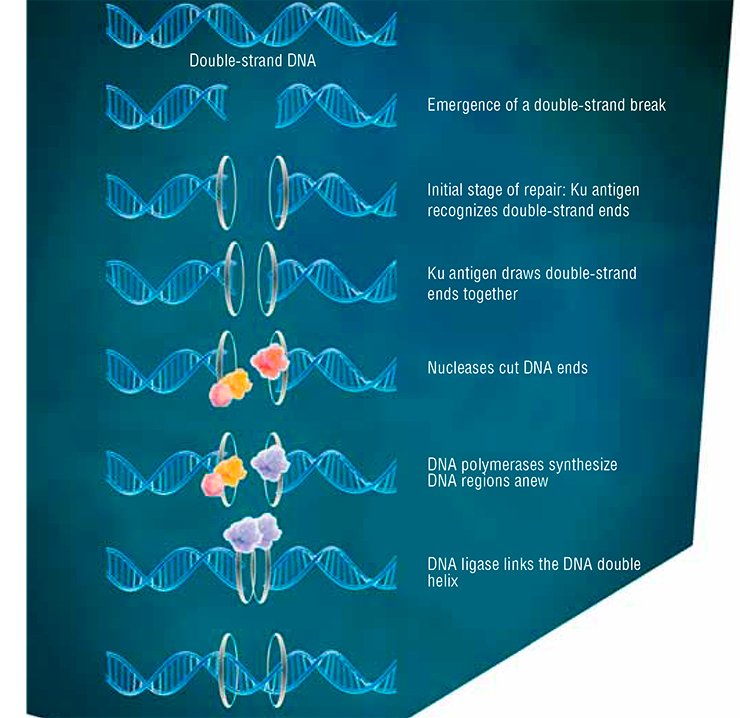
In the other phases of the cell cycle, DNA is repaired by nonhomologous end joining, implemented by a specialized protein ensemble, with Ku antigen playing “first fiddle” (Davis et al., 2013). This way of repair is less precise, but, paradoxically, it is prevalent in the cells of higher eukaryotes, which also include humans.
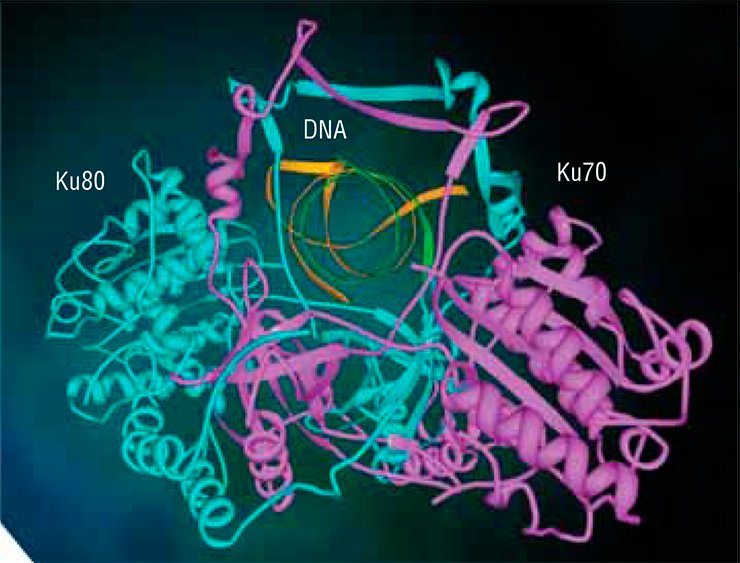
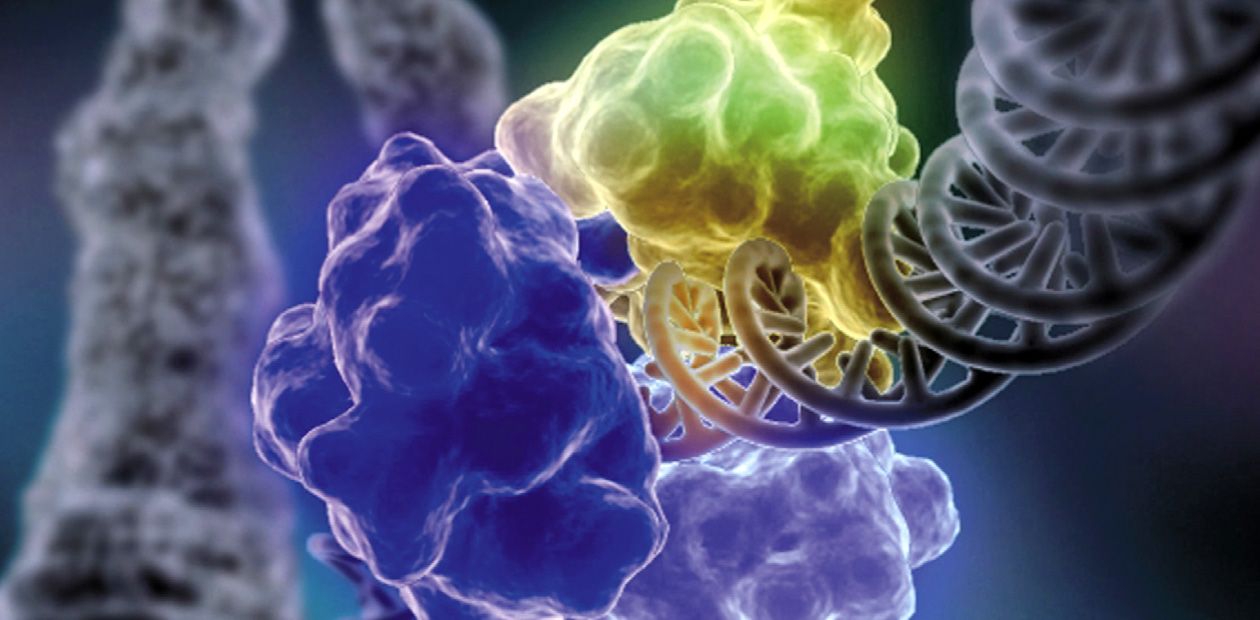
The Ku antigen is involved in the first stage of repair, when it recognizes the ends of double-strand breaks, brings them together, and holds them in this way until the initial DNA structure is restored. The structure of the Ku antigen itself is perfectly adapted for such an activity: this molecule is shaped as a padlock with a massive body and thin shackle or as a wicker basket with a handle (Walker et al., 2001). Two molecules of the Ku antigen slipped on the ends of a DNA molecule interact with one another like two halves of a magnetic fastener that secures a string of beads around the neck.
The “bead-on-string” type of binding between Ku antigen and DNA allows the protein to slide easily along DNA, which is necessary for the repair protein complex to work. The basket-like folding of Ku antigen allows for a relatively free access of other repair proteins to DNA in its “handle” part, while the body of the basket serves as a platform for assembling repair proteins around the lesion.
A universal approach
However, the importance of Ku antigen for the body is not confined to its role in DNA repair. This explains why the protein in question was discovered several times when different processes were studied.
For the first time, Ku antigen was discovered in the patients with autoimmune diseases (polymyositis, sclerodermitis, and systemic lupus erythematosus), as a protein antigen that induces antibody production. The discovered protein got its name fr om the first letters in the surname of the patient in whom such antibodies were first detected. Later, the antibodies to Ku antigen were also found in the patients with other autoimmune diseases, in particular, rheumatoid arthritis.
At first, Ku antigen was found to be located mostly in cell nuclei; then its ability to interact with DNA was discovered. It took several years to ascertain the fact that this protein is involved in the repair of DNA double-strand breaks. After this discovery, the data on the role of Ku antigen in various cellular processes started coming as if from the horn of plenty.
Note that in some cases the role of Ku antigen is determined by its direct involvement in DNA repair. This refers to a unique physiological phenomenon that takes place at the early stages of the development of the immune system cells and which provides the body with the so-called innate cell-mediated immunity.
The matter is that in the immune cell precursors the gene region that encodes the variable part of antibodies (or receptors) and is responsible for antigen recognition consists of several repeated DNA segments belonging to three classes—V, D, and J. During the immune cell differentiation, the so-called V(D)J recombination produces a population of cells carrying different combinations of V, D, and J segments (one from each class), which makes the cells able to recognize various antigens. These events, yielding the primary diversity of antigens (receptors), take place well ahead of the first encounters of an organism with bacteria, viruses, and any other real pathogens. Since recombination implies the emergence of double-strand breaks in DNA, Ku antigen provides for their repair. Correspondingly, any impairment in the work of this protein leads to a severe combined immunodeficiency.
Thus, wise Mother Nature resolves seemingly unrelated problems, using one and the same “device”.
A part-time employee
In addition to the activity directly associated with the repair of DNA double-strand breaks, Ku antigen is an important player in the cellular processes that, one way or another, maintain the DNA structure.
In particular, a decrease in the Ku antigen content in the cell negatively influences replication, i.e. synthesis of a daughter DNA molecule; this protein also contributes to chromatin looping in the nucleus. Interestingly, despite the fact that no DNA double-strand breaks are produced in these two processes, they include a mild “melting” (formation of unpaired DNA regions) of the double helix.
Ku antigen also contributes to maintaining the integrity of telomeres, terminal chromosome regions with a specific structure of repeated sequences protected by specialized proteins. On the one hand, telomeres prevent the chromosome ends from being recognized as a double-strand break that has to be repaired. On the other hand, in normal cells telomeres become shorter with each next division. In this sense, telomeres serve as a certain clock that specifies the lifespan of a cell. Note that telomeres in cancer cells have a constant length, and these cells retain the ability to divide.
The above examples demonstrate the role of Ku antigen in maintaining and implementing genetic information, but the amazing capabilities of this protein are even greater. For example, it has been quite unexpectedly discovered that Ku antigen is present not only in the nucleus, but also in the cell cytoplasm, wh ere it functions independently of DNA.
As is known, Ku antigen comprises two separate subunits. The Ku70 subunit, located in the cytoplasm, is involved in the cell response to toxic impacts or, more precisely, in regulating apoptosis, i. e. genetically programmed cell death in the case of its damage.
Here, the effect of Ku70 may be dual. On the one hand, this subunit can bind the cytoplasmic protein Bax, which contributes to the mitochondrial apoptotic pathway. In this case cell suicide is triggered by Bax relocation into mitochondria, while Ku70 prevents apoptosis by blocking this protein.
On the other hand, Ku70 is able to free Bax from the attached molecules of ubiquitin, a small cellular protein that acts as a kind of “black spot”. Thus, Ku70 sustains the life of a Bax molecule “condemned to death” by assisting in triggering apoptosis.
Note that many cancer cells are able to “escape” the suicide program, which underlines the necessity to focus on the Ku antigen activity in tumor cells.
However, there are other things that come as a surprise. Ku antigen appears to be also present on the outer cell surface, wh ere it is bound to the cytoplasm membrane. However amazing it may seem, there Ku antigen acts as a “fifth column,” helping some bacteria and viruses to enter the cell (Martinez et al., 2005; Munakata et al., 2005).
In addition, Ku antigen is responsible for adhesion of cells to one another and to the intercellular matrix, the totality of extracellular proteins that supports the structure of biological tissue (Muller et al., 2005). The Ku antigen on the outer side of the membrane is also involved in cell migration, helping a specialized protease (an enzyme that destroys proteins) to “gnaw through” a way for the cell by partially destroying the intercellular matrix (Muller et al., 2005). This last “talent” of Ku antigen may do a poor service to the body affected by cancer by enhancing metastatic growth.
Too much water drowned the miller
The diversity of Ku antigen functions suggests that both the increased and decreased levels of this protein may be significant factors that influence cancer cell activity.
For example, multiple myeloma (a kind of tumor) cells contain a truncated variant of the normal Ku80 subunit, Ku80v. This polypeptide is able to interact normally with the other subunit (Ku70), and the resulting dimer, with DNA double-strand ends. However, this “flawed” variant of Ku antigen is unable to bind the catalytic subunit of DNA-dependent protein kinase, thereby arresting the initial stages of DNA double-strand break repair.
On the contrary, the content of normal Ku antigen variant is elevated in the cells of another tumor, B-cell chronic lymphoid leukemia, which intensifies DNA repair. As a result, the cancer cells become more resistant to ionizing radiation and other genotoxic agents, which makes antitumor radiotherapy inefficient. These examples illustrate the necessity to determine precisely the Ku antigen status in cells. Several approaches are currently available for this purpose.
As is known, the level of expression—the conversion of hereditary information of a gene into a functional product, RNA or protein—is usually assessed in clinical and research laboratories with the help of real-time polymerase chain reaction (PCR). In PCR, the messenger RNA (“how-to” description for synthesizing a protein) is extracted from cells and used as a template for “reverse” synthesis of the corresponding DNA sequence. The resulting DNA can be then repeatedly copied.
Scientists have devised and implemented several smart ways of such DNA “propagation”; however, even the most sophisticated method demonstrates only the presence or absence of the target DNA sequence (and, thus, of the corresponding protein) in the analyzed sample. It is rather difficult to estimate the quantity of an active protein in this way. Besides, it is impossible to find out whether a truncated variant, Ku80v, is present: this variant is synthesized from the same RNA template; only after that the “extra” fragment is cleaved.
In addition to PCR, to assess protein levels several varieties of enzyme immunoassay are used; the analysis uses antibodies that interact with a certain region of the target protein and requires two types of antibodies—primary and secondary.
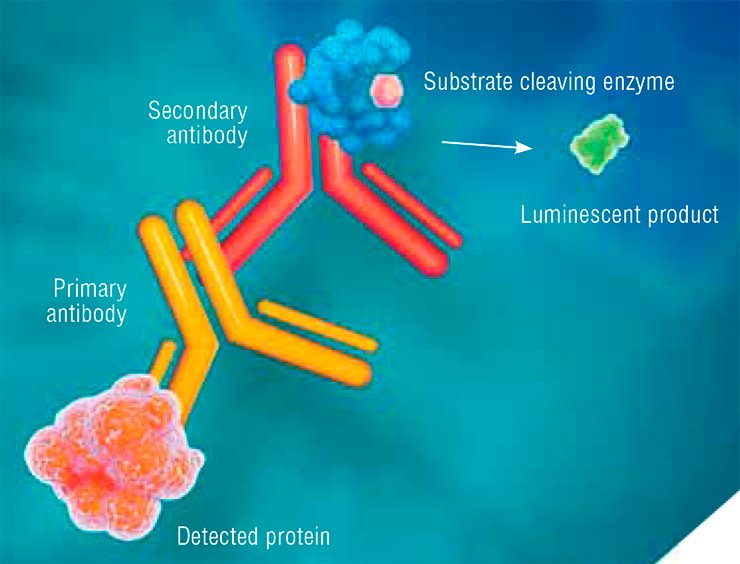
Primary antibodies are produced in laboratory animals (most frequently, rabbits or mice) in response to the injection of a foreign protein. Secondary antibodies, intended for recognizing the primary ones, are produced in another animal species. An enzyme able to cleave a certain substrate and thus produce a luminescent product is attached to such secondary antibodies.
Thus, the in vitro enzyme immunoassay implies a chain of interactions between the target protein, primary antibody, and secondary antibody. The result is luminescence, which helps to detect the target protein.
Since the antibodies interact with a particular region of the protein sequence, this method makes it possible to distinguish between the Ku80 subunits of different lengths. Two types of primary antibodies are necessary for this purpose: one type will bind to the region present in both variants of the polypeptide and the second one, only to the region present in the full-sized variant.
On the job
All the above mentioned methods can detect Ku antigen either indirectly (PCR) or separately from its cellular functions. The research team of the Laboratory of Bioorganic Chemistry of Enzymes with the Institute of Chemical Biology and Fundamental Medicine (Siberian Branch, Russian Academy of Sciences, Novosibirsk, Russia) has elaborated a fundamentally different assay for Ku antigen directly on the “jobsite,” i.e. on the DNA molecule (Ilina et al., 2010).
This assay uses the ability of Ku antigen to interact with the so-called AP sites, i.e. the DNA regions that lost their nitrogenous bases. These “lacunas” in the DNA “text” are among the most prevalent lesions. Ku antigen can bind to the DNA containing AP sites, thereby protecting them from cleavage (Ilina et al., 2008). However, when a double-strand break is located close to AP sites, Ku antigen cleaves them, thereby contributing to the “cleanup” of break ends and preparing the break for repair (Roberts et al., 2010).
The new method for detecting Ku antigen, designed by the Novosibirsk team, uses the fluorescently or radioactively labeled DNA probe with an AP site. Having bound to such DNA, Ku antigen forms an unstable covalent complex with it. A special treatment irreversibly crosslinks Ku antigen to DNA, which in this case acts as a tag. The label on the DNA probe makes it possible to record the resulting products reliably and with high sensitivity, and even determine their quantity.

The major advantage of this approach is its ability to detect all active species of Ku antigen that can bind to DNA, including the truncated variant. The lower cost and simpler performance as compared with the standard enzyme immunoassay are also appealing.
Ku antigen is the key protein underlying the activity of the system repairing DNA double-strand breaks which emerge at a high rate during tumor radiotherapy. In this sense, the level of Ku antigen may be regarded as an important prognostic factor when the efficiency and purposefulness of this treatment are assessed.
The new assay for Ku antigen, proposed by Novosibirsk researchers, makes it possible to assess the level of active protein in cancer cells and may be used for making prognoses along with the currently used PCR assays.
Research into Ku antigen properties is in progress; now it is focused on the interactions of this protein not with single lesions, but mainly with complex clustered damage (Kosova et al., 2014). We hope that the nearest future will bring a deeper insight into the properties of this amazing protein as well as the possibility of its application.
References
Khodyreva S. N., Lavrik O. I. How cells repair DNA // Science First Hand. 2007. No. 15. P. 82—89.
Davis A. J., Chen D. J. DNA double strand break repair via non-homologous end-joining // Transl. Cancer Res. 2013. V. 2. P. 130–143.
Ilina E. S., Lavrik O. I., Khodyreva S. N. Ku antigen interacts with abasic sites // Biochem. Biophys. Acta. 2008. V. 1784. P. 1777–1785.
Ilina E. S., Khodyreva S. N., Berezhnoy A.E., et al. Tracking Ku antigen levels in cell extracts with DNA containing abasic sites // Mutat. Res. 2010. V. 685. P. 90–96.
Walker J. R., Corpina R. A., Goldberg J. Structure of the Ku heterodimer bound to DNA and its implications for double-strand break repair // Nature. 2001. V. 412. P. 607–614.
This work was supported by the RFBR (project # 13-04-01426) and by the programme Molecular and Cell Biology, Russian Academy of Sciences