Life Began with RNA
Why does a paper about RNA, not other, more well-known organic molecules such as DNA or proteins appear in the cycle of articles on the origin of life? Perhaps our readers have heard about RNA, but what? Most likely, nothing noteworthy for the simple reason that so far only experts in biology know that RNA consists of ‘magic’molecules that formed the background for the origin of life. Once in ancient times, on the just cooled Earth, there arose and existed a mysterious ‘RNA world’…
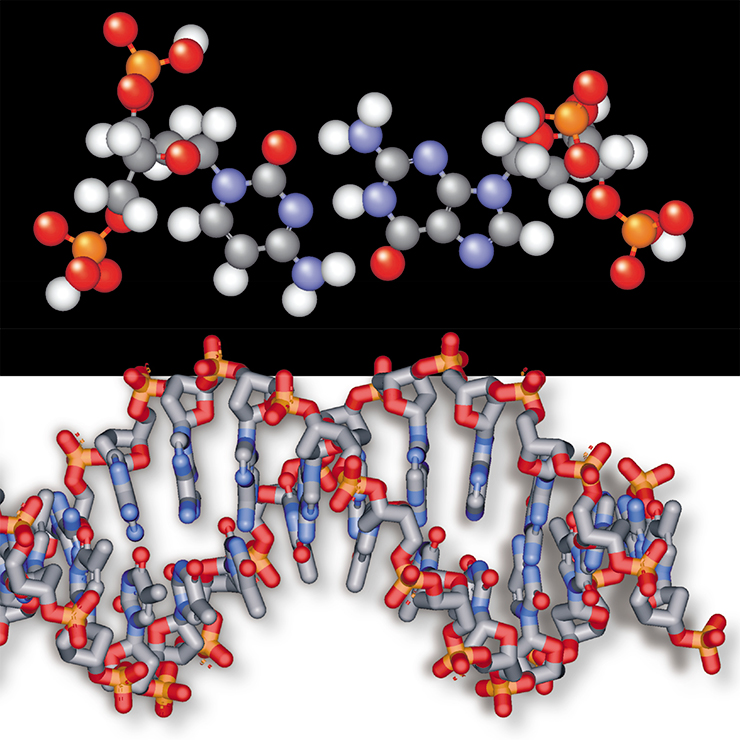
Before embarking on our journey to the ‘beginning of the beginning’, let us stock up with necessary knowledge about the structure of nucleic acids – DNA (deoxyribonucleic) and RNA (ribonucleic). In its chemical structure, RNA is a twin, although not an identical twin, of DNA, the major repository of genetic information in a living cell. Nucleic acids are polymeric macromolecules composed of separate elements, nucleotides. Molecules of a five-carbon sugar linked by phosphates form the backbone of these macromolecules. One nitrogen base is attached to each sugar molecule. The nucleotides, whose only distinction from one another is different nitrogen bases, are designated by letters A, U, G, and C (in RNA) and A, T, G, and C (in DNA).
RNA is somewhere in between – it is only a carrier of information from DNA. Subsequently, discoveries followed, leading to a radically different view of RNA
The main difference between these two types of nucleic acids is in their carbon component. The sugar of RNA is ribose; of DNA, deoxyribose: DNA has a hydrogen atom (H) at the site where RNA has an oxy group (OH). What seem to an inexperienced eye to be insignificant differences have striking results. For example, DNA exists mainly in the form of the well-known rigid helixes, where two DNA strands are held together by hydrogen bonds formed between complementary nucleotides.
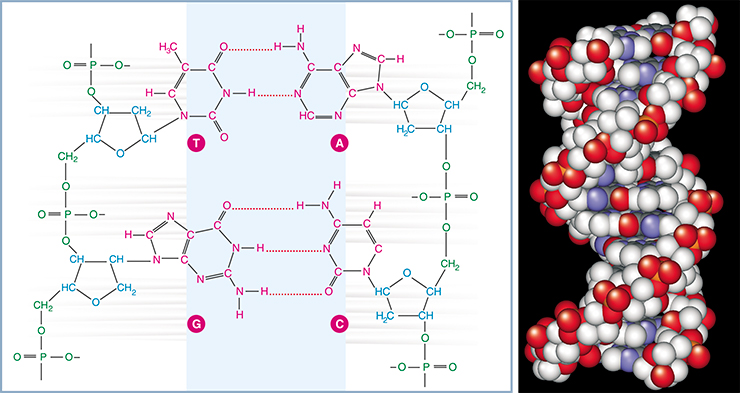
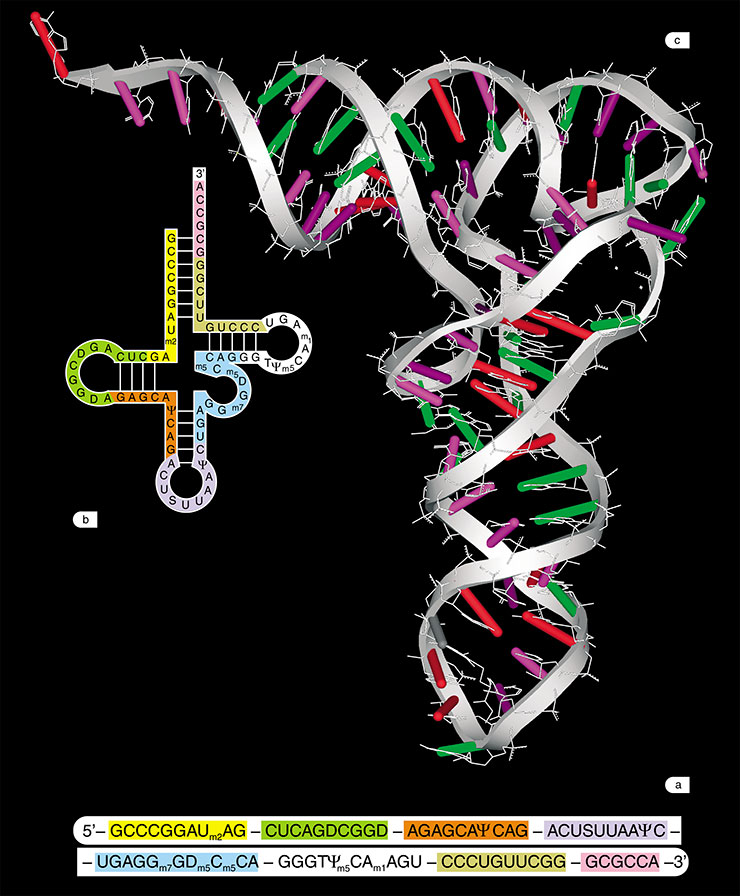
RNA is also capable of forming double-stranded helices similar to DNA helices; however, RNA in the majority of cases exists as intricate tangle structures. These structures are formed not only due to the mentioned hydrogen bonds, but also because of the ribose oxy group, which is able to form additional hydrogen bonds and interact with the phosphoric acid and metal ions. Globular RNA structures not only outwardly resemble protein structures, but also approximate proteins in their features – they are capable of interacting with various molecules, both small and polymeric.
What Can be Considered ‘Alive’?
Why do we call RNA the foremother of currently existing life? To answer this question, let us determine where the border is between the living and the inanimate.
Since scientists from various fields are involved in attempts to solve the problem of the origin of life, each of them operates with the terms of his own science. Chemists will certainly remember the term ‘catalyst’; mathematicians will think of the word ‘information’. Biologists will consider a system alive only if it contains a substance (genetic program) capable of duplicating (or, more simply speaking, of reproducing). It is also necessary that certain changes in hereditary information emerge during such copying, thereby bringing into being new variants of systems – that is, this implies the existence of evolution. Moreover, biologists will definitely note that such systems must be isolated spatially. Otherwise, the evolved, more progressive systems will be unable to make use of their advantages, since their more efficient catalysts and other products float away into the ambient medium.
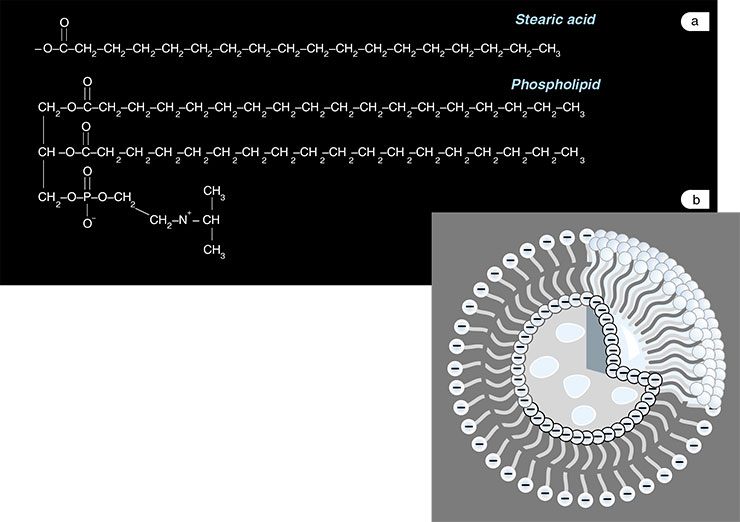
Thus, how were the first molecular systems separated from the environment? For example, colonies of molecules could keep together due to adsorption on some mineral surface or dust particles. However, the most primitive systems might as well possess a genuine membrane envelope, similar to modern living cells. The point is that such a ‘protocell’ with lipid membrane could form very easily. Many molecules with charged groups (for example, fatty acids) in an aqueous medium form microscopic vesicles, or liposomes. This word is undoubtedly known well to half of our readers, since liposomes are widely used in cosmetics – miniature fatty capsules are stuffed with vitamins and other biologically active substances. However, what was the filling of ancient ‘protocells’? It turns out that RNA lays claim to this role of ‘stuffing’.
Can RNA do Everything?
Undoubtedly, life should have commenced from the formation of ‘skilled’ molecules that would be able to reproduce themselves and keep all the other ‘household duties’ necessary for existence of the cell. However, neither DNA nor proteins are suitable to the role of such skilled workers. DNA is a perfect repository of genetic information, but is unable to reproduce itself. Proteins are unsurpassed catalysts but cannot function as ‘genetic programs’. Thus, we meet the paradox of the chicken and the egg: DNA cannot be formed without the protein; the protein cannot be formed without DNA. And here it becomes apparent that RNA can do everything. Nevertheless, we will not run off ahead.
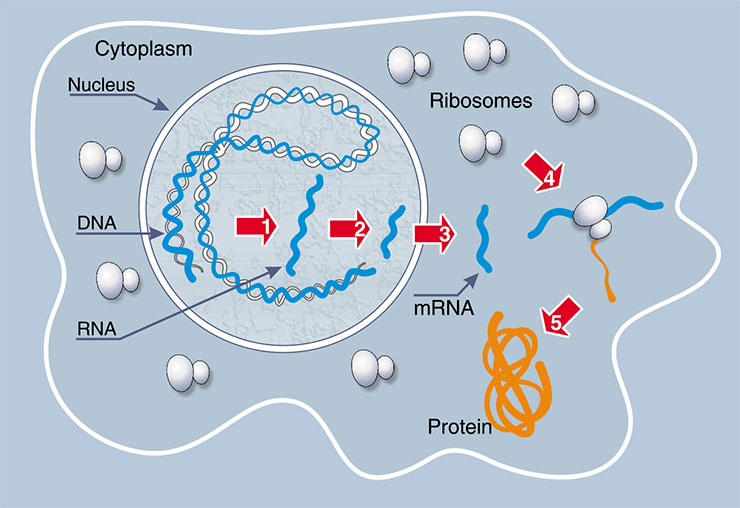
First, consider the well-known RNA functions related to the work (expression) of a gene in a cell. When the gene is switched on, DNA is unwound locally to allow for synthesizing an RNA copy of the genetic program. A complex processing of the RNA copy by specialized proteins produces the messenger RNA (mRNA) – the program for synthesis of the corresponding protein. This RNA is transported from the cell nucleus to the cytoplasm to bind to specialized cell structures – ribosomes – molecular ‘plants’ for protein synthesis. The protein is synthesized from activated amino acids attached to specialized transfer RNA (tRNA); moreover, each amino acid is attached to its own specific tRNA. Thanks to tRNA, the amino acid is fixed in the catalytic center of the ribosome, where it is ‘stitched’ to the protein chain synthesized. It is clear from this chain of events that RNA molecules play the key role in the decoding of the genetic information and in the biosynthesis of protein.
As researchers delved more deeply into various biosynthetic processes, they frequently began discovering previously unknown RNA functions. For example, in addition to the process of transcription (RNA synthesis through copying a DNA region), DNA can sometimes be synthesized from an RNA template. This process, named reverse transcription, is involved in development cycles of numerous viruses, including the notorious oncogenic viruses and HIV-1, causing AIDS.
Thus, the flow of genetic information is not unidirectional – from DNA to RNA – as it was considered before. The role of DNA as the original, major carrier of genetic information now causes certain doubts, even more so since numerous viruses (influenza virus, tick-borne encephalitis virus, and many others) do not use DNA as a carrier of genetic information at all. Their genome is built exclusively on RNA. Then, discoveries poured out one after another, and they made us look at RNA from a different standpoint.
A Jack-of-All-’Molecules’
 The most amazing thing was the discovery of RNA’s catalytic ability. It was previously generally believed that only proteins and enzymes are capable of catalyzing reactions. For example, scientists for a long time were unable to isolate the enzymes involved in cleaving and ligating certain RNA. It appeared after long studies that RNA perfectly coped with the task on its own. The RNA structures acting like enzymes were named ribozymes (by analogy with enzymes, protein catalysts). A variety of numerous ribozymes were soon discovered. Viruses and other simple infectious agents use ribozymes particularly widely for handling their RNA. Thus, RNA proved to be a Jack-of-all-trades: it is able to carry hereditary information, act as a catalyst, act as a transport machinery for amino acids, and form highly specific complexes with proteins.
The most amazing thing was the discovery of RNA’s catalytic ability. It was previously generally believed that only proteins and enzymes are capable of catalyzing reactions. For example, scientists for a long time were unable to isolate the enzymes involved in cleaving and ligating certain RNA. It appeared after long studies that RNA perfectly coped with the task on its own. The RNA structures acting like enzymes were named ribozymes (by analogy with enzymes, protein catalysts). A variety of numerous ribozymes were soon discovered. Viruses and other simple infectious agents use ribozymes particularly widely for handling their RNA. Thus, RNA proved to be a Jack-of-all-trades: it is able to carry hereditary information, act as a catalyst, act as a transport machinery for amino acids, and form highly specific complexes with proteins.
It was an x-ray structure assay revealing details of the structure of ribosome crystals that gave definitive certainty as to the actual existence of the ‘RNA world’. Researchers expected to find proteins catalyzing the stitching of amino acids into a protein sequence. How astonished they were to find no protein structures at all in the catalytic center of ribosomes — it was solely built of RNA! Thus, RNA molecules perform all the key stages of protein biosynthesis. The discussion was settled as to the potential existence of the ‘RNA world’ as a specific stage of biological evolution. Indeed, the complete picture has yet to be reconstructed, as many problems remain unsolved. For example, specialized protein enzymes activate amino acids in the modern cell and attach them to the corresponding tRNA. It is reasonable to ask here whether this reaction could be realized without proteins, only with involvement of RNA, and whether RNA itself could catalyze synthesis of RNA from nucleotides or attachment of nitrogen bases to sugar. Once ribozymes were discovered, these potential capabilities of RNA did not cause much doubt. However, science requires that hypotheses be experimentally verified.
Darwinian Evolution in a Tube
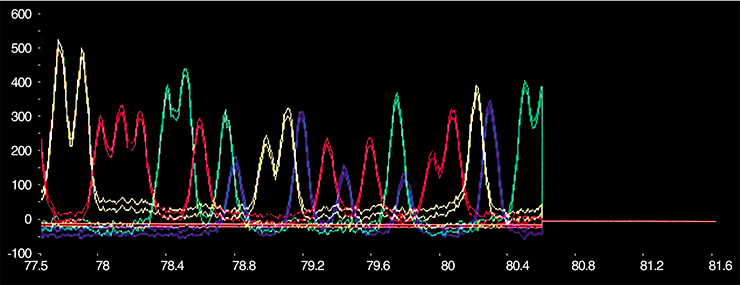
Good method often brings about a revolution in science. An illustrative example of such a method is polymerase chain reaction (PCR), allowing nucleic acids to be reproduced in unlimited amounts. Its essence is as follows. To reproduce DNA, PCR employs the enzymes DNA polymerases – that is, the same enzymes that synthesize complementary DNA strands from activated monomers, nucleotides, during cell reproduction.
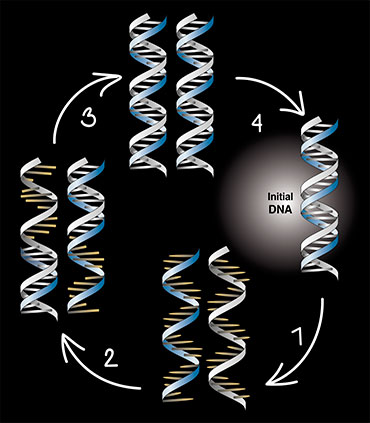
Thus, a mixture of activated nucleotides, DNA polymerase, and the so-called primers – oligonucleotides complementary to the ends of DNA to be multiplied – is placed in the tube with the target DNA. When a DNA solution is heated, the DNA strands untwist. During the subsequent cooling, primers bind to DNA strands, forming short fragments of helix structures. The enzyme attaches nucleotides to primers and synthesizes the strand complementary to the initial DNA strand. The reaction makes two DNA strands from one. Another repeat, and we have four strands, and n repeats produce 2n DNA molecules. It’s all very simple.
The advent of PCR and development of the methods for DNA chemical synthesis formed the background for a tremendous technology of molecular selection. Its essence is simple too: a multitude of molecules displaying various properties (the so-called molecular library) is synthesized to select from this mixture those molecules that display the desirable characteristics.
Libraries of nucleic acids are mixtures of molecules with the same length but differing in the nucleotide sequence. It is possible to obtain these when all four nucleotides are added simultaneously at each stage of nucleotide sequence elongation during chemical synthesis in an automated sequencer. Each of the nucleotides is incorporated in the growing nucleic acid with equal probability; thus, four variants of sequences will be generated at each stage. If a nucleotide sequence with a length of n nucleotides is thus synthesized, the number of various molecules will amount to 4n. Since fragments with a length of 30–60 monomers are conventionally used, 430 to 460 different molecules are generated! These orders of magnitude are customary, perhaps, only for astronomers.
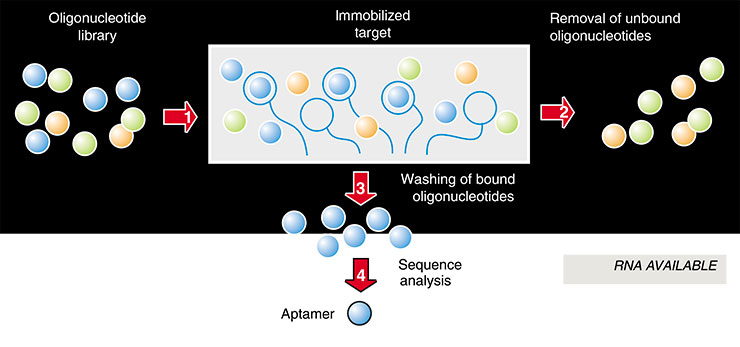
As nucleic acids fold into different spatial structures depending on their composition, an analysis of statistical sequences gives a tremendous diversity of molecules differing in their properties. RNA is transcribed from the formed DNA by the enzyme RNA polymerase. Consequently, a library of single-stranded RNA is thus formed. The next procedure is selection: the RNA solution is passed through a column containing an insoluble carrier with chemically attached target molecules to ‘fish out’ the so-called future aptamer, that is, the RNA capable of binding particular molecules. Then the column is rinsed to remove the unbound RNA, and then the RNA retarded in the column due to binding to the target molecules is washed (for example, by heating the column).
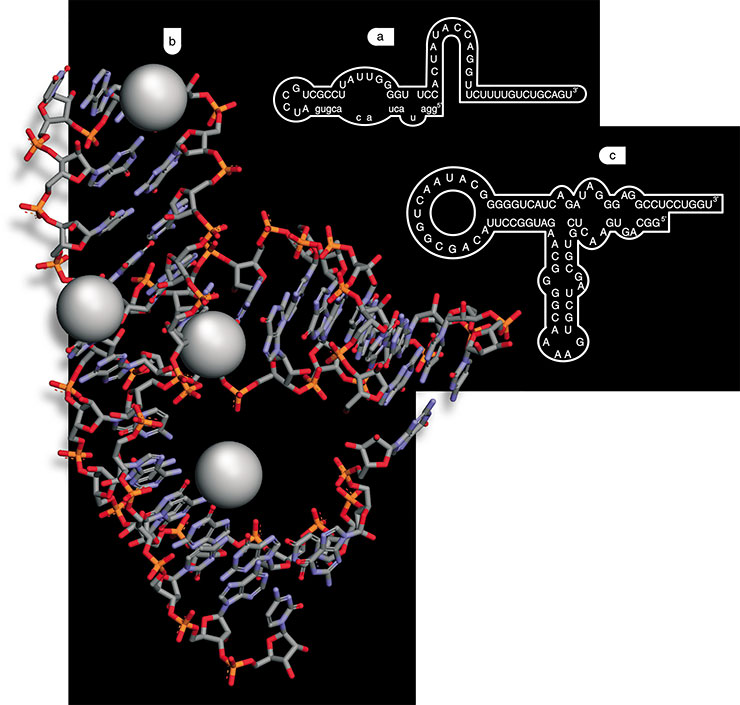
DNA copies are produced from the selected RNA to obtain from them common double-stranded DNA molecules. The target RNA aptamers are generated from the latter, and now they are ready for reproduction by PCR in an unlimited quantity. Indeed, this is an ideal situation; the practical realization is more complex. Usually, the initial RNA preparation contains an immense excess of ‘foreign molecules’, which are difficult to remove. Consequently, the RNA obtained is repeatedly passed through the column to separate the RNA forming the most stable complexes with the target molecules. Thousands of manifold RNA aptamers capable of binding specifically to various organic compounds and molecules have been obtained using this method.
This scheme of molecular selection is applicable to designing molecules with any properties. For example, RNA capable of catalyzing reactions of RNA and protein syntheses – attachment of nitrogen bases to ribose, polymerization of activated nucleotides on the RNA strand, and attachment of amino acids to RNA – have been obtained. These studies confirmed once again that RNA molecules with specific structures and functions could have originated from random polymers under conditions of prebiotic evolution.
RNA Available
The technology of molecular selection has a high potential. This tool makes it possible to search for the necessary targets even if initially one has no idea what the structure of the target molecules should be. However, once the selection procedure is developed and designed, the necessary molecules can be separated based on the required properties, only then dealing with the question of how the required properties are achieved. Below is an example of isolation of the RNA capable of binding to cell membranes and modulating its permeability.
The ancient ribocytes (RNA cells) needed to consume ‘nutrients’ from the environment, excrete metabolic products, and divide during their reproduction. All these processes require control over membrane permeability. We assume that ribocytes contained no other functional molecules except for RNA; consequently, certain RNA molecules should possess the ability to interact with membranes. However, from the chemical standpoint, they are absolutely unsuitable for the role of membrane permeability regulators.
The membranes of modern cells and liposomes, composed of fatty acids, are negatively charged. Since RNA is also charged negatively, Coulomb’s law forces them to be repelled from the lipid surface; even more so, they cannot penetrate into the lipid layer. The only known way in which nucleic acids interact with the membrane surface is via divalent metal ions. These positively charged ions may play the role of bridges between the negatively charged groups on the membrane surface and the phosphate groups of the nucleic acid. These bridge interactions are rather weak; hence, only a very large nucleic acid through a large number of weak interactions is capable of binding to the membrane surface. In such a way, the Lilliputians tied Gulliver to the ground with a multitude of thin strings.
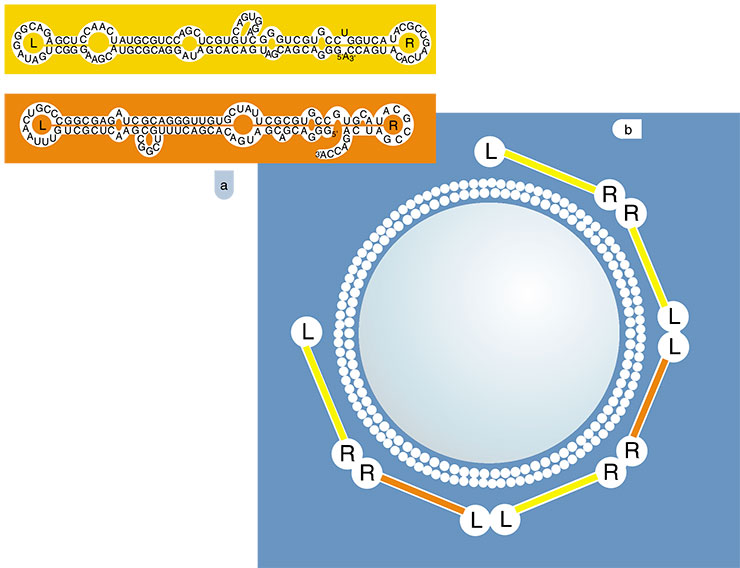
Here is a case where molecular selection was of great help. Several RNA molecules were successfully isolated from the RNA library that not only successfully bound to membranes, but with a high enough RNA concentration even tore them! These RNA molecules displayed quite unusual properties – it was as if they helped one another: the mixture of various types of molecules bound to membranes more readily than the molecules of one type. A study of the secondary structure of these RNA molecules made the situation clear. It appeared that they had loops with complementary regions. Due to these regions, the ‘membrane’ RNAs can form complexes–communities capable of building numerous contacts with the membrane, thereby performing what is beyond the power of one RNA molecule.
This selection experiment suggests that RNA had an additional resource for acquiring new features via forming intricate supramolecular complexes. This mechanism could have been also used for confinement of evolving RNA systems as colonies on surfaces even before these systems developed their own isolating membranes.
The ‘RNA World’: Existed, Exists, and Will Exist!
Numerous data demonstrate that the ‘RNA world’ actually existed. However, it is not clear where. Some specialists believe that the initial stages of evolution occurred not on Earth, that already functional systems were delivered to our planet and adapted to the local conditions. Anyway, this does not change the essence of the matter from both a chemical and biological standpoint. However, the remaining enigma is what particular processes in the environment brought ribocytes into being and what components maintained their existence. Indeed, nucleotides, required for the life of ribocytes, are complex molecules. It is difficult to imagine how such substances could be formed under conditions of prebiotic synthesis.

Altogether, it is not improbable that the ancient RNA differed essentially from modern molecules. Unfortunately, it is impossible to detect experimentally traces of this ancient RNA, since the question concerns a time separated from us by billions of years. Even the rocks of that age have long ago disintegrated into sand. Thus, the only possibility is to model experimentally the processes that could occur at the earliest stages of molecular evolution.
Why did there occur a transition from the ‘RNA world’ into the modern world? Proteins, possessing a considerably greater set of chemical groups than RNA, are better catalysts and structural elements. Presumably, some ancient RNA commenced using protein molecules as ‘implements’. Such RNA, which was in addition able to synthesize useful molecules for its purposes from environmental elements, acquired advantages in reproduction. The corresponding aptamers and ribozymes were naturally selected. And then the evolution did its work: the translation machinery emerged and the responsibility for catalysis gradually was handed over to proteins. These implements appeared so convenient that dislodged their ‘hosts’ from many former spheres of their activities.
The reader has the right to ask: Why it is necessary at all to study the evolution of RNA? After all, the ancient ‘RNA world’ has long ago disappeared. Is it just art for art’s sake and satisfaction of the interests of fanatics of science? Indeed, the present is incomprehensible without knowledge of the past. Research into the evolution and potentials of RNA may prompt new directions in the search for the processes occurring in modern living cells. For example, powerful systems have recently been discovered that regulate activities of genes with involvement of double-stranded RNAs; the cell uses these systems for protection against viral infections. This ancient system of cell defense will most likely soon find application in medicine.
Thus, it is not surprising that research in nucleic acids continues to be one of the ‘hot spots’ in molecular biology. Thanks to its unique properties, RNA has ever expanding applications in medicine and technology. The ‘RNA world’, which originated in time immemorial, will not only continue its invisible existence in our cells, but also will be regenerated in new biotechnologies.
The Editorial Board thanks Drs. V. V. Koval, S. D. Myzina, and A. A. Bondar (Institute of Chemical Biology and Fundamental Medicine, Siberian Branch of the Russian Academy of Sciences) for their assistance in preparation of this article